validated biomarkers
evidence-graded insights
peer-reviewed studies
test options to choose from
Everything you need to choose and use the right test
How each test works
Full biomarker listings
Side-by-side comparison
Clinical decision support
Real-world case studies
Workflow and ordering
Ready to explore the range?
Download the complete product guide for a detailed look at every test, or speak with one of our Clinical Application Specialists to talk through which option is right for your practice.
Validated biomarkers
Evidence-graded insights
Panel from one sample
Day turnaround
Ready to start referring?
Download the Essentials spec sheet for the complete picture, or speak with a Clinical Application Specialist about whether Essentials is right for your patients.
Validated biomarkers
Evidence-graded insights
Panels from one sample
Pathogen targets
Ready to start referring?
Download the Comprehensive spec sheet for the complete picture, or speak with a Clinical Application Specialist about whether Comprehensive is right for your patients.
validated biomarkers
evidence-graded insights
peer-reviewed studies
test options to choose from
Ready to start referring?
Download the Extended spec sheet for the complete picture, or speak with a Clinical Application Specialist about whether Extended is right for your patients.
The gut microbiome is
a measurable biological system
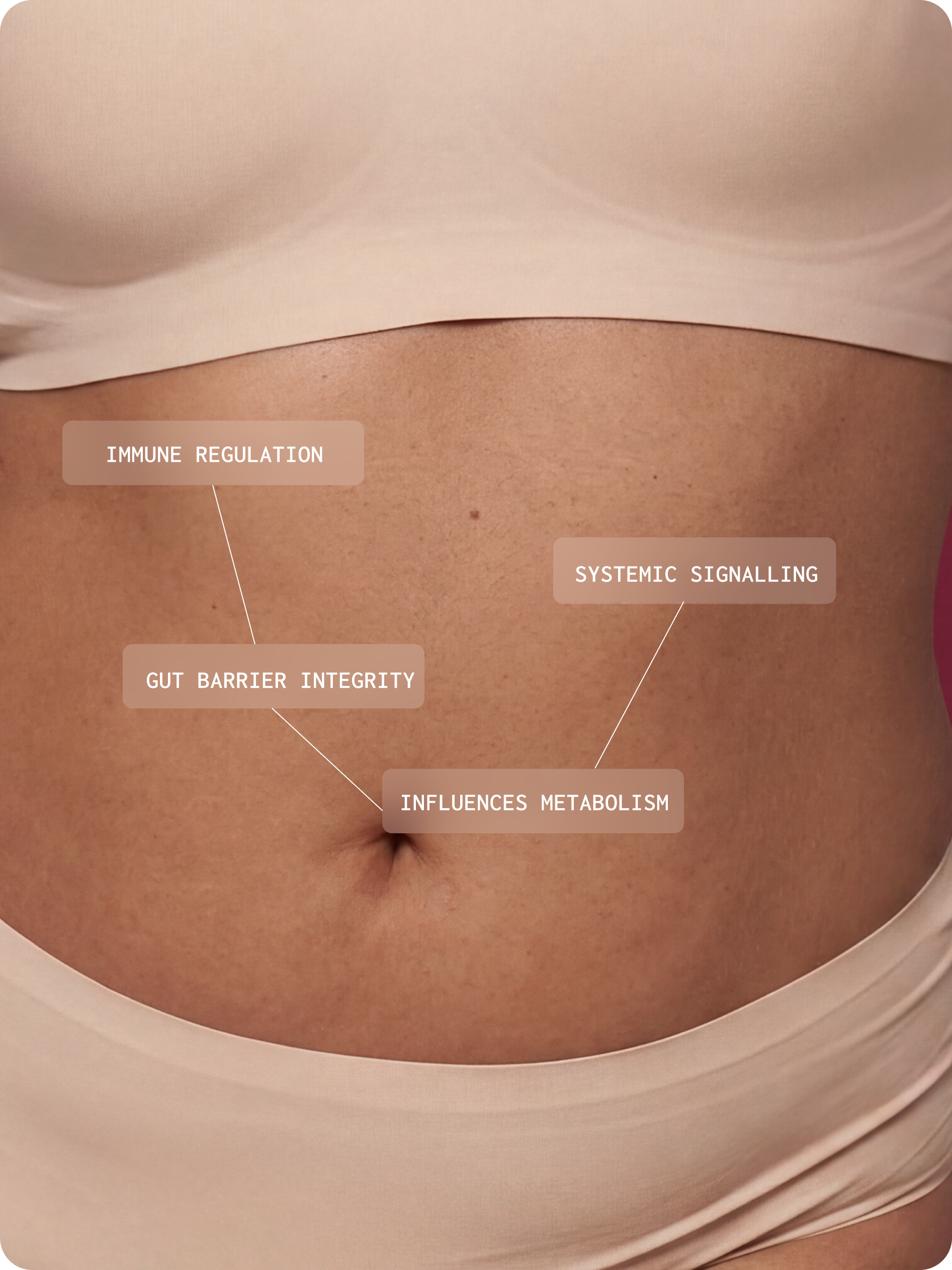
The gut microbiome is a dynamic biological system that is essential to normal human physiology, influencing immune regulation, gut barrier integrity, metabolism, and systemic signaling.1
It is a clinically relevant biological system with meaningful implications for complex and multi-system presentations.
The healthy function of the microbiome depends on a balanced ecosystem
The nature of these interactions is not fixed, it depends on the gut environment and overall ecological balance. These interactions generally fall into four categories
Bacteria
Archaea
Viruses
Fungi
Microbial interactions are shaped by balance, not fixed labels
The nature of these interactions is not fixed, it depends on the gut environment and overall ecological balance. These interactions generally fall into four categories:
Present without measurable benefit or harm to the host.
Contributes positively to metabolism, immune regulation or barrier integrity.
Typically neutral, but may cause harm when ecological balance shifts.
Capable of causing disease or disrupting normal function.
Microbes convert fuel into compounds that influence health
Gut microbes feed on available substrates including dietary fibre, proteins and host-derived compounds and convert them into metabolites that interact with body systems.1,2 What they produce depends on which microbes are present, the functional pathways they carry, and the fuel sources available to them.
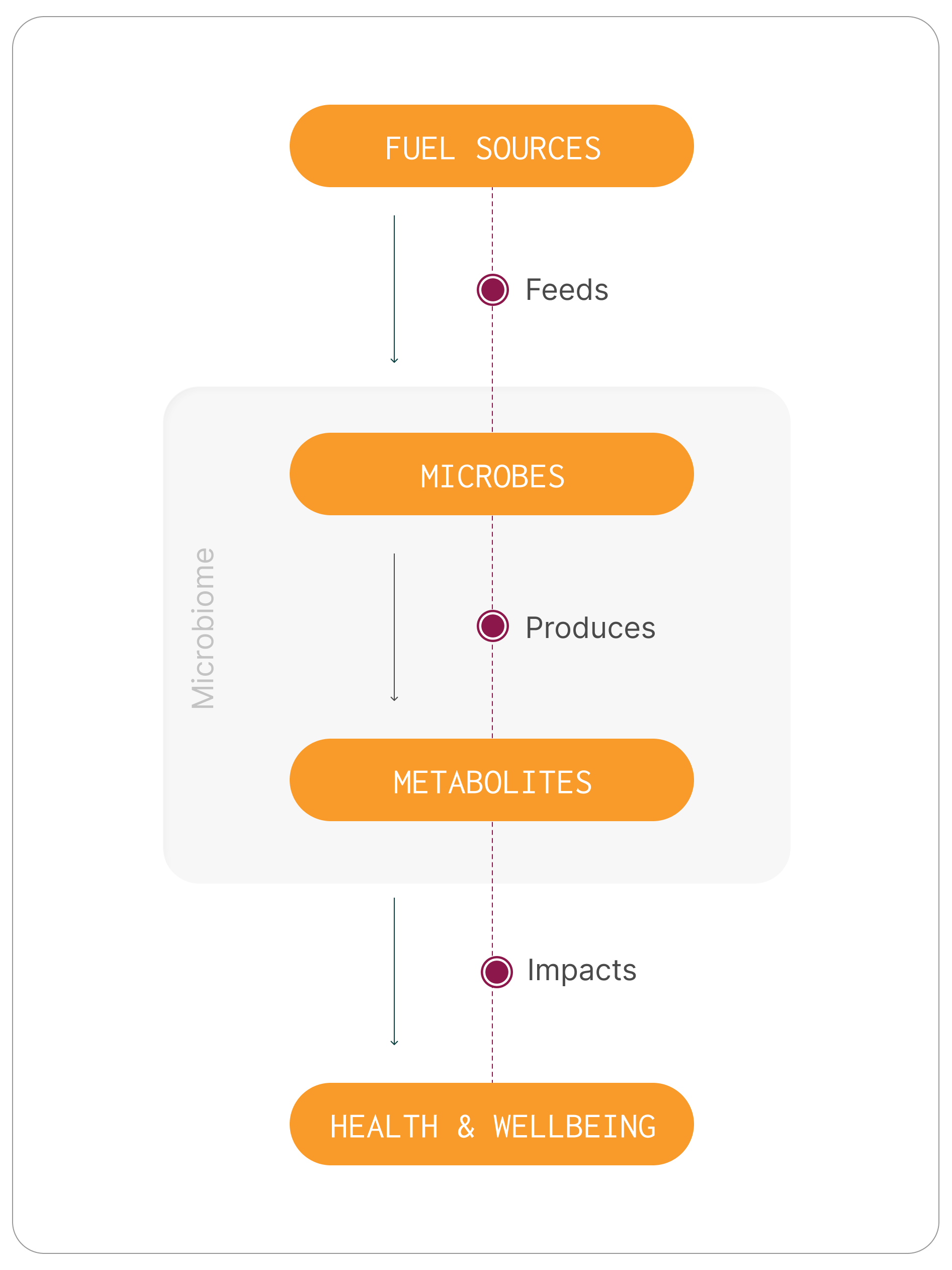
Key helpful functions of the gut microbiome
Microbes influence human physiology through the compounds they produce and the signals they generate. These microbial metabolites interact with immune pathways, support gut barrier integrity, and contribute to broader metabolic regulation.
Metabolism
Microbial metabolites influence energy use, lipid processing and systemic metabolic balance
Immune regulation
The gut microbiome helps regulate immune responses and maintain immune balance
Gut barrier function
Microbial activity supports epithelial integrity
Gut–brain & systemic signalling
Microbial activity supports epithelial integrity
The microbiome functions as part of human biology — not separate from it.
It’s not enough to know which microbes are present, you need to know what the ecosystem is doing
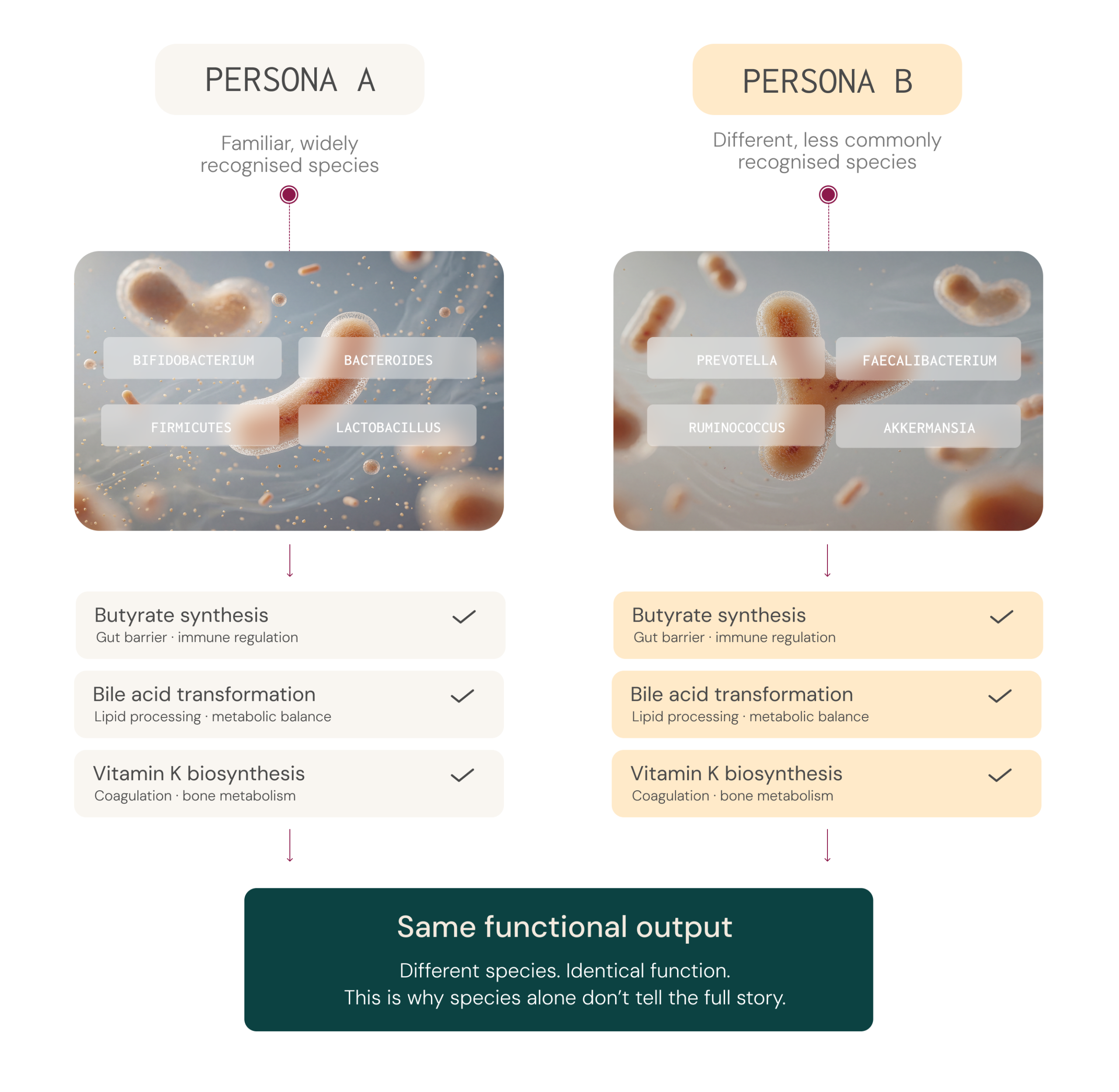
Microbiome species vary widely between individuals, yet different species can produce the same essential metabolites. Two people with very different microbial profiles may share comparable functional output.4-6
This is why clinical insight increasingly focuses on metabolic activity, such as butyrate production, rather than species detection alone..

Microbiome insight adds a systems-level lens to clinical care.
The microbiome interacts with multiple biological systems simultaneously, it can provide additional interpretive insight in chronic and complex cases.
In multi-system presentations not fully explained by traditional markers, understanding microbial function adds a broader systems-level perspective.
In multi-system presentations not fully explained by traditional markers, understanding microbial function adds a broader systems-level perspective.
In multi-system presentations not fully explained by traditional markers, understanding microbial function adds a broader systems-level perspective.
Ecological balance
Functional pathways
Metabolic output
Patient context
Key Takeaway
The human microbiome is a measurable biological system that functions as part of normal human physiology.
Community-wide measures such as diversity and functional output can be more clinically meaningful than detection of specific species.
As understanding of the microbiome grows, it is becoming an increasingly valuable part of clinical assessment, especially in chronic or multi-system presentations.
References:
- Sender R. Fuchs S. & Milo R. Revised Estimates for the Number of Human and Bacteria Cells in the Body.PLoSBiology 14 e1002533 (2016).
- HuttenhowerC. et al. Structure function and diversity of the healthy human microbiome. Nature 486 207–214 (2012).
- Jiang Y. Che L. & Li S. C. Deciphering the personalized functional redundancy hierarchy in the gut microbiome. Microbiome 14 17 (2025).
- Fan Y. & Pedersen O. Gut microbiota in human metabolic health and disease. Nat. Rev.Microbiol. 19 55–71 (2021).
- Lynch S. V. & Pedersen O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375 2369–2379 (2016).
- Koh A. DeVadderF. Kovatcheva-Datchary P. & Bäckhed F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 165 1332–1345 (2016).
- ZhernakovaA. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352 565–569 (2016).
- De Filippis F. et al. Distinct genetic and functional traits of human intestinalPrevotellacopri strains are associated with different habitual diets. Cell Host Microbe 25 444–453 (2019).
- Zhang Z. J. et al.Comprehensive analyses of a large human gutBacteroidales culture collection reveal species- and strain-level diversity and evolution. Cell Host Microbe 32 1853–1867 (2024).
- Tian L. et al. Deciphering functional redundancy in the human microbiome. Nat Commun 11 6217 (2020).
- denBestenG. et al. The role of short-chain fatty acids in the interplay between diet gut microbiota and host energy metabolism. Journal of Lipid Research 54 2325–2340 (2013).
- Morrison D. J. & Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 7 189–200 (2016).
- Venkatesh M. et al. Symbiotic Bacterial Metabolites Regulate Gastrointestinal Barrier Function via the Xenobiotic Sensor PXR and Toll-like Receptor 4. Immunity 41 296–310 (2014).
- Honda K. & Littman D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535 75–84 (2016).
- Furusawa Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504 446–450 (2013).
- Arpaia N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504 451 (2013).
- MukhopadhyaI. & Louis P. Gut microbiota-derived short-chain fatty acids and their role in human health and disease. Nat Rev Microbiol 23 635–651 (2025).
- Singh V. et al. Butyrate producers “The Sentinel of Gut”: Their intestinal significance with and beyond butyrate and prospective use as microbial therapeutics. Front.Microbiol. 13 (2023).
- Ren T. et al. Indole Propionic Acid Regulates Gut Immunity: Mechanisms of Metabolite-Driven Immunomodulation and Barrier Integrity. JMicrobiolBiotechnol 35 e2503045 (2025).
- Cryan J. F. et al. The Microbiota-Gut-Brain Axis. Physiological Reviews 99 1877–2013 (2019).
- Matsuura M. Structural Modifications of Bacterial Lipopolysaccharide that Facilitate Gram-Negative Bacteria Evasion of Host Innate Immunity. Front. Immunol. 4 (2013).
- ZamyatinaA. & Heine H. Lipopolysaccharide Recognition in the Crossroads of TLR4 and Caspase-4/11 Mediated Inflammatory Pathways. Front. Immunol. 11 (2020).
- Zhu M. et al. C-reactive protein and cancer risk: a pan-cancer study. BMC Med 20 301 (2022).
- RidkerP. M. et al. Relationship of C-reactive protein reduction to cardiovascular event reduction following treatment with canakinumab. The Lancet 391 319–328 (2018).
- Pradhan A. D. Manson J. E. Rifai N. Buring J. E. &RidkerP. M. C-Reactive Protein Interleukin 6 and Risk of Developing Type 2 Diabetes Mellitus. JAMA 286 327–334 (2001).
- d’HennezelE. Abubucker S. Murphy L. O. & Cullen T. W. Total Lipopolysaccharide from the Human Gut Microbiome Silences Toll-Like Receptor Signaling. mSystems 2 e00046-17 (2017).
- Khorsand B. et al. Overrepresentation of Enterobacteriaceae and Escherichia coli is the major gut microbiome signature in Crohn’s disease and ulcerative colitis. Front. Cell. Infect.Microbiol. 12 (2022).
- Thompson K. N. et al. Alterations in the gut microbiome implicate key taxa and metabolic pathways across inflammatory arthritis phenotypes. Science Translational Medicine 15 eabn4722 (2023).
- Desai M. S. et al. A Dietary Fiber-Deprived Gut Microbiota Degrades the Colonic Mucus Barrier and Enhances Pathogen Susceptibility. Cell 167 1339-1353.e21 (2016).
- Zheng J. et al.Noninvasivemicrobiome-based diagnosis of inflammatory bowel disease. Nat Med 30 3555–3567 (2024).
- Qin J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490 55 (2012).
- Tuomainen M. et al. Associations of serumindolepropionicacid a gut microbiota metabolite with type 2 diabetes and low-grade inflammation in high-risk individuals. Nutrition & Diabetes 8 35 (2018).
- Peron G. et al. A Polyphenol-Rich Diet Increases the Gut Microbiota Metabolite Indole 3-Propionic Acid in Older Adults with Preserved Kidney Function. Molecular Nutrition & Food Research 66 2100349 (2022).
- SpraggeF. et al. Microbiome diversity protects against pathogens by nutrient blocking. Science 382 eadj3502 (2023).
- ByndlossM. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357 570–575 (2017).
The gut microbiome is
a measurable biological system
The gut microbiome is fundamental to normal human physiology. It is a dynamic ecosystem that influences immune regulation, gut barrier integrity, metabolism, and systemic signalling.⁴
This is a two-way relationship. The microbiome contributes to metabolic activity, produces bioactive compounds, and participates in immune signalling.
In turn, the body provides nutrients that support gut microbes and influences how they behave. Diet, immune activity, and gut conditions all shape which microbes thrive and what they do.4,5
The microbiome is an clinically relevant biological system with meaningful implications for complex and multi-system presentations. Clinically, this is especially important for patients whose symptoms don’t fit neatly into a single diagnosis or one clear cause.

The gut microbiome is a diverse microbial community
The gut microbiome is a diverse community of microorganisms that actively shape gut function through metabolism, competition, and interactions with the immune system and gut lining. This community includes:
Bacteria

Archaea

Viruses

Fungi
The role of gut microbes depends on ecological context
The role a microbe plays is not fixed, it depends on the gut environment and the overall ecological balance of the community. Host-microbe interactions generally fall into four broad categories:
Present without measurable benefit or harm to the host.
Contributes positively to metabolism, immune regulation or barrier integrity.
Typically neutral, but may cause harm when ecological balance shifts.
Capable of causing disease or disrupting normal function.
Microbes convert fuel into compounds that influence health
Gut microbes feed on available substrates, including dietary fibre, proteins, and host-derived compounds, and convert them into metabolites that interact with body systems.4,6
Some of these metabolites act locally within the gut, influencing barrier integrity and immune signalling. Others enter the bloodstream and affect organs and processes throughout the body.
What is produced depends on:
Species: The microbes present
Functional pathways: The key metabolic functions they perform
Dominant substrates: The main fuel sources they utilise
Gut environment: Conditions present in the gut such as pH, oxygen availability, and nutrient availability.

This is why microbiome assessment increasingly focuses on functional capacity and metabolic output — not just species detection.
It’s not enough to know which microbes are present you need to know what the ecosystem is doing
Microbiome composition varies significantly between individuals.⁷ While certain microbial groups are commonly found across populations, the relative abundance, diversity, and functional output differ from person to person.
Function matters more than labels
Microbiome composition varies significantly between individuals.⁷ This variability is shaped by many factors, including diet, medication use, genetics, environmental exposures, and lifestyle, all of which influence both microbial composition and metabolic capacity.⁷
Two individuals can share similar broad microbial groups yet differ considerably in their specific species or strains,² and those species and strains may carry different genes with different metabolic capacities.8,9 The reverse is also true: two people with very different microbial profiles may share comparable functional outputs, because different species can produce the same metabolites.2,3,10
Key helpful functions of the gut microbiome
The gut microbiome influences health not by controlling organs directly, but through the metabolites it produces and the signals they generate.
While the microbiome’s influence on the body is broad, four well-characterised areas are particularly relevant to clinical practice: metabolism, immune regulation, gut barrier integrity, and systemic signalling. Importantly, this relationship is bidirectional – the body’s environment, including diet, immune activity, and gut conditions, also shapes the microbiome.4,5
Metabolism
Gut microbes act like a metabolic organ, producing thousands of different metabolites from dietary and host-derived compounds.11,12 Some of these metabolites stay in the gut, where they can affect inflammation, transit time, and the gut lining.
Others cross into the bloodstream and can influence inflammation, energy balance, lipid processing, and metabolic regulation throughout the body.11,12
Immune regulation
The microbiome continuously interacts with immune cells within the gut-associated lymphoid tissue (GALT).14 Microbial antigens and metabolites help calibrate immune responsiveness – supporting tolerance to harmless signals while maintaining the ability to respond to genuine threats. For example, some short-chain fatty acids promote the production of regulatory T cells that help prevent over-reactive inflammation.15,16 Rather than simply stimulating immunity, the microbiome plays a central role in immune homeostasis.14
Gut barrier function
The gut barrier is a set of layers that separates gut contents from the bloodstream: a mucus layer, a single layer of epithelial cells, and tight junctions between those cells. Microbial metabolites — including butyrate and some tryptophan-derived indoles – can strengthen this barrier by providing an energy source to gut cells and promoting tight junction integrity.6,13,17,19
Thus, barrier function is not purely structural — it is actively maintained by microbial metabolism.
Systemic signalling
Microbial metabolites interact with the body’s receptors and influence signalling pathways beyond the gut, including enteroendocrine signalling, neural circuits, and immune pathways.4,20 The gut and brain communicate through nerves (including the vagus nerve), hormones, and immune signals. Gut microbes can influence these pathways by producing or modifying neuroactive compounds and by modulating inflammation levels.20
The microbiome functions as part of human biology — not separate from it.
What shapes a person's gut microbiome
The gut microbiome is not static, it is continually adjusting to its environment. A range of modifiable and non-modifiable factors influence which microbes thrive and what they do, many of which are modifiable.⁷
Diet
Medications
Infections and inflammation
Lifestyle and environment
What happens when the microbiome is out of balance
There is no single "perfect" microbiome. Healthy people can have very different microbial profiles. However, when the balance shifts significantly, it can affect what the microbiome produces and how the body responds. An imbalanced microbiome — sometimes called dysbiosis — often involves one or more of the following:
Loss of helpful microbes (for example, fibre-fermenting short-chain fatty acid producers)
Overgrowth of potentially harmful microbes (sometimes called pathobionts)
Reduced diversity (fewer different types of microbes)
A shift in what the microbiome is doing — such as producing fewer protective molecules and more inflammatory molecules
How imbalances can affect the immune system and inflammation
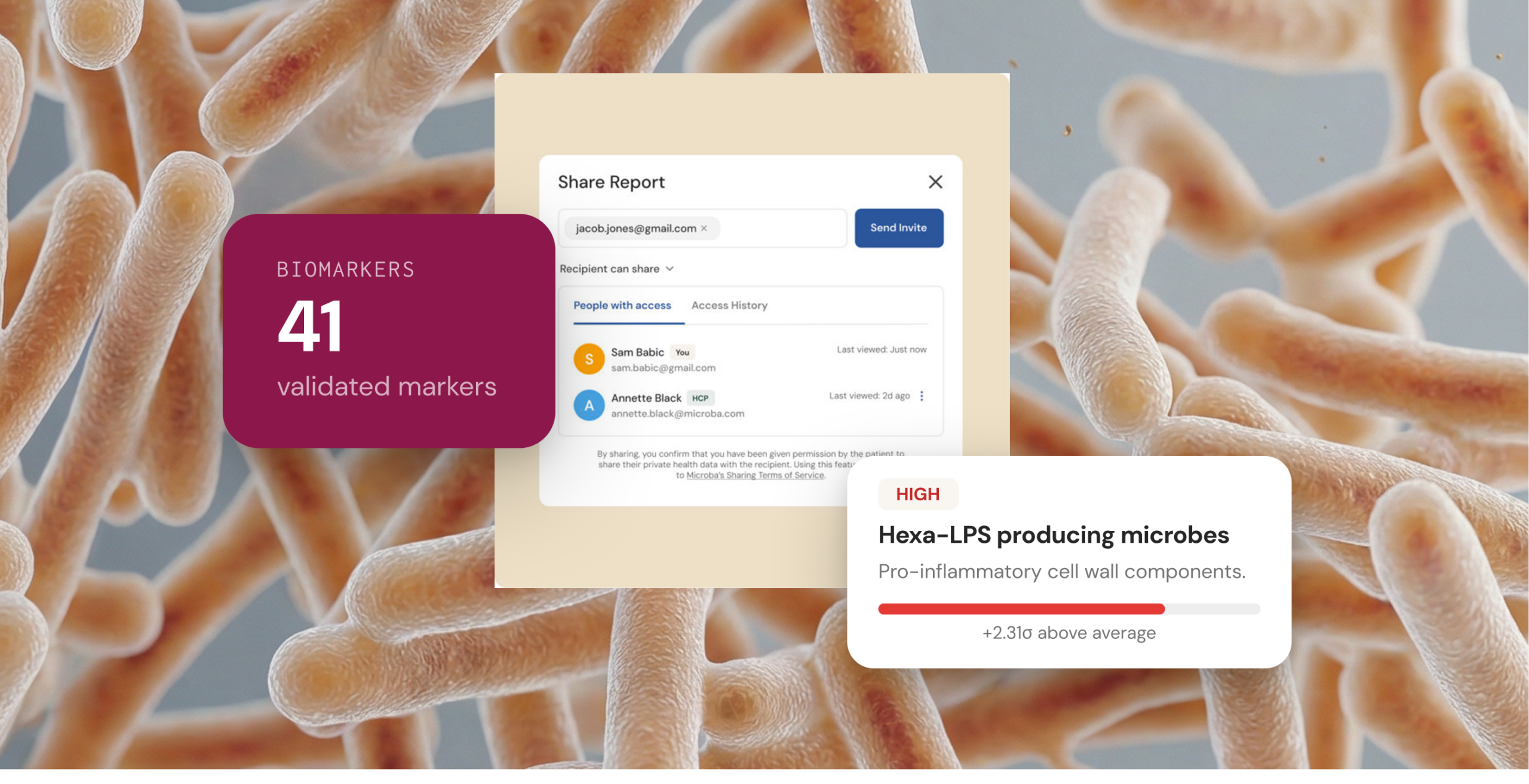
The immune system is one of the most direct ways the gut microbiome can affect whole-body health. When the balance of microbial metabolites shifts, with fewer anti-inflammatory products such as butyrate15,16 and more pro-inflammatory products such as hexa-acylated LPS21,22 – immune cells may stay activated for prolonged periods, leading to low-grade chronic inflammation. This type of inflammation is linked with many chronic diseases.23-25
Pro-inflammatory LPS (endotoxin)
Specific types of LPS (hexa-acylated or higher) are strong triggers for inflammation because they can activate a receptor on immune cells called TLR4, which can initiate an inflammatory cascade.21 Other types of LPS are weaker and may even help block this signalling.26 When the balance shifts toward bacteria that produce strongly inflammatory LPS, inflammatory signalling can increase. Increased levels of these bacteria have been observed in Crohn’s disease, rheumatoid arthritis, and ankylosing spondylitis.27-28
Mucus layer damage when fibre is low
If dietary fibre is limited, some microbes can switch to using mucus as a fuel source. In animal studies, a fibre-deprived diet increased mucus-degrading bacteria, thinned the mucus barrier, and increased susceptibility to infection and inflammation.29 In humans, the relative abundance of mucin-degrading pathways has been positively correlated with faecal calprotectin, a clinical marker of inflammation in the gut.7
Lower butyrate and IPA-producing species
Butyrate is a short-chain fatty acid that serves as the primary energy source for colon cells and plays a key role in immune regulation. Indole-propionic acid (IPA) is a tryptophan-derived metabolite also involved in immune regulation and maintaining intestinal barrier integrity. Reduced levels of butyrate-producing species and plasma IPA have been associated with inflammatory bowel diseases and type 2 diabetes.30-32
Defending against pathogens
A diverse microbiome with adequate levels of butyrate producers can help prevent overgrowth of harmful microbes by competing for space and nutrients, producing antimicrobial substances, and maintaining key signalling pathways.34-35 When the microbiome is disrupted, for example after antibiotics, this protective function can be compromised, leaving people more susceptible to pathogen infections.
What matters most in clinical interpretation
Because the microbiome interacts with multiple biological systems, it can provide additional interpretive context in chronic and complex presentations -- particularly when traditional markers do not fully explain a patient's symptoms. The microbiome is not best understood through a "good vs. bad bugs" lens. As outlined in the preceding sections, clinical interpretation is more informative when it considers these key factors: the ecological balance of the community, the functional pathways present, the metabolic output of those pathways, and the individual patient's context.
Ecological balance: community stability and diversity
Functional pathways: what microbes do, not just which are present
Equally, two similar-looking profiles may differ in what they actually produce. Assessing what the microbiome is capable of doing often provides more clinically meaningful insight than cataloguing which species are present.
Metabolic output: the compounds produced and their effects
Patient context: immune status, diet, and environment
Key Takeaway
The gut microbiome is a measurable, modifiable part of human physiology. It functions as an ecosystem — and like any ecosystem, its health depends on the balance, diversity, and functional capacity of the community as a whole, not just the presence or absence of individual organisms.
For clinicians, the most useful insight from the microbiome is often functional rather than taxonomic: what the microbial community is producing, how that interacts with the patient’s immune system, gut barrier, and metabolic regulation, and how those outputs may be shifting in the context of that individual’s diet, medications, and clinical picture.
For clinicians, microbiome insights provide an additional lens for understanding systemic regulation, particularly in chronic and complex symptom patterns.
As our understanding of the microbiome deepens, so does the opportunity to use it as a meaningful part of clinical assessment, particularly for patients with chronic or multi-system presentations where traditional markers tell only part of the story.

Explore our full resource Hub
Access evidence-based guides, webinars, clinical tools, and educational resources designed to help you deepen your understanding of the gut microbiome and support confident clinical conversations.
References:
- Sender R. Fuchs S. & Milo R. Revised Estimates for the Number of Human and Bacteria Cells in the Body.PLoSBiology 14 e1002533 (2016).
- HuttenhowerC. et al. Structure function and diversity of the healthy human microbiome. Nature 486 207–214 (2012).
- Jiang Y. Che L. & Li S. C. Deciphering the personalized functional redundancy hierarchy in the gut microbiome. Microbiome 14 17 (2025).
- Fan Y. & Pedersen O. Gut microbiota in human metabolic health and disease. Nat. Rev.Microbiol. 19 55–71 (2021).
- Lynch S. V. & Pedersen O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375 2369–2379 (2016).
- Koh A. DeVadderF. Kovatcheva-Datchary P. & Bäckhed F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 165 1332–1345 (2016).
- ZhernakovaA. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352 565–569 (2016).
- De Filippis F. et al. Distinct genetic and functional traits of human intestinalPrevotellacopri strains are associated with different habitual diets. Cell Host Microbe 25 444–453 (2019).
- Zhang Z. J. et al.Comprehensive analyses of a large human gutBacteroidales culture collection reveal species- and strain-level diversity and evolution. Cell Host Microbe 32 1853–1867 (2024).
- Tian L. et al. Deciphering functional redundancy in the human microbiome. Nat Commun 11 6217 (2020).
- denBestenG. et al. The role of short-chain fatty acids in the interplay between diet gut microbiota and host energy metabolism. Journal of Lipid Research 54 2325–2340 (2013).
- Morrison D. J. & Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 7 189–200 (2016).
- Venkatesh M. et al. Symbiotic Bacterial Metabolites Regulate Gastrointestinal Barrier Function via the Xenobiotic Sensor PXR and Toll-like Receptor 4. Immunity 41 296–310 (2014).
- Honda K. & Littman D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535 75–84 (2016).
- Furusawa Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504 446–450 (2013).
- Arpaia N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504 451 (2013).
- MukhopadhyaI. & Louis P. Gut microbiota-derived short-chain fatty acids and their role in human health and disease. Nat Rev Microbiol 23 635–651 (2025).
- Singh V. et al. Butyrate producers “The Sentinel of Gut”: Their intestinal significance with and beyond butyrate and prospective use as microbial therapeutics. Front.Microbiol. 13 (2023).
- Ren T. et al. Indole Propionic Acid Regulates Gut Immunity: Mechanisms of Metabolite-Driven Immunomodulation and Barrier Integrity. JMicrobiolBiotechnol 35 e2503045 (2025).
- Cryan J. F. et al. The Microbiota-Gut-Brain Axis. Physiological Reviews 99 1877–2013 (2019).
- Matsuura M. Structural Modifications of Bacterial Lipopolysaccharide that Facilitate Gram-Negative Bacteria Evasion of Host Innate Immunity. Front. Immunol. 4 (2013).
- ZamyatinaA. & Heine H. Lipopolysaccharide Recognition in the Crossroads of TLR4 and Caspase-4/11 Mediated Inflammatory Pathways. Front. Immunol. 11 (2020).
- Zhu M. et al. C-reactive protein and cancer risk: a pan-cancer study. BMC Med 20 301 (2022).
- RidkerP. M. et al. Relationship of C-reactive protein reduction to cardiovascular event reduction following treatment with canakinumab. The Lancet 391 319–328 (2018).
- Pradhan A. D. Manson J. E. Rifai N. Buring J. E. &RidkerP. M. C-Reactive Protein Interleukin 6 and Risk of Developing Type 2 Diabetes Mellitus. JAMA 286 327–334 (2001).
- d’HennezelE. Abubucker S. Murphy L. O. & Cullen T. W. Total Lipopolysaccharide from the Human Gut Microbiome Silences Toll-Like Receptor Signaling. mSystems 2 e00046-17 (2017).
- Khorsand B. et al. Overrepresentation of Enterobacteriaceae and Escherichia coli is the major gut microbiome signature in Crohn’s disease and ulcerative colitis. Front. Cell. Infect.Microbiol. 12 (2022).
- Thompson K. N. et al. Alterations in the gut microbiome implicate key taxa and metabolic pathways across inflammatory arthritis phenotypes. Science Translational Medicine 15 eabn4722 (2023).
- Desai M. S. et al. A Dietary Fiber-Deprived Gut Microbiota Degrades the Colonic Mucus Barrier and Enhances Pathogen Susceptibility. Cell 167 1339-1353.e21 (2016).
- Zheng J. et al.Noninvasivemicrobiome-based diagnosis of inflammatory bowel disease. Nat Med 30 3555–3567 (2024).
- Qin J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490 55 (2012).
- Tuomainen M. et al. Associations of serumindolepropionicacid a gut microbiota metabolite with type 2 diabetes and low-grade inflammation in high-risk individuals. Nutrition & Diabetes 8 35 (2018).
- Peron G. et al. A Polyphenol-Rich Diet Increases the Gut Microbiota Metabolite Indole 3-Propionic Acid in Older Adults with Preserved Kidney Function. Molecular Nutrition & Food Research 66 2100349 (2022).
- SpraggeF. et al. Microbiome diversity protects against pathogens by nutrient blocking. Science 382 eadj3502 (2023).
- ByndlossM. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357 570–575 (2017).
Microbiome research is no longer about detection, it’s about function
Over the past two decades, microbiome research has moved from simply identifying which microbes are present to understanding what they do and how they interact with the host.2 Large population studies have shown that while microbial species differ widely between individuals, many core metabolic functions are preserved.3
EARLY 2000S
Species cataloguing and taxonomic classification
2010’s
Functional profiling and metagenomic sequencing
current
Clinical integration and longitudinal intervention studies
Advances in metagenomic sequencing now enable functional profiling — assessing microbial gene content and pathway capacity rather than relying on species-level associations alone.3 Longitudinal studies and controlled human interventions continue to clarify how the microbiome responds to diet, medication and environmental exposure.4,5
Microbial function significantly
influences chronic conditions
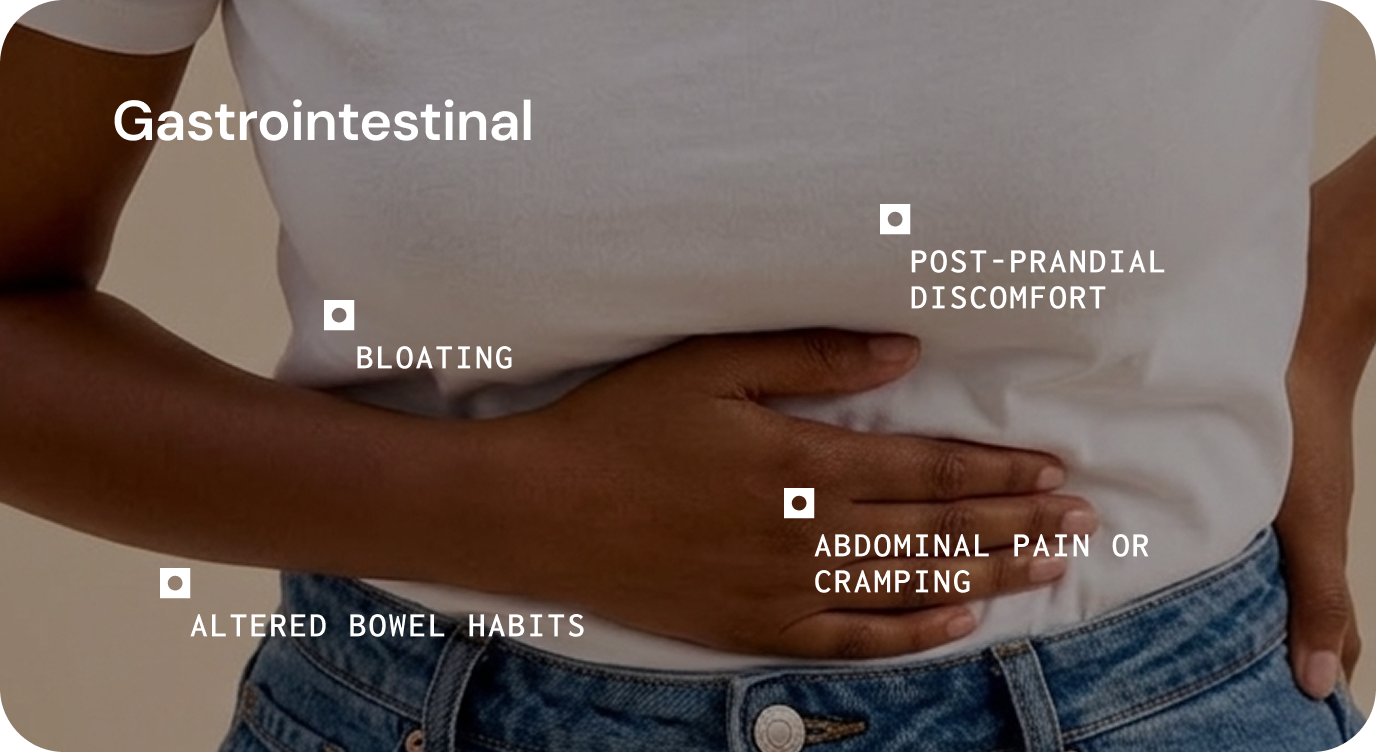
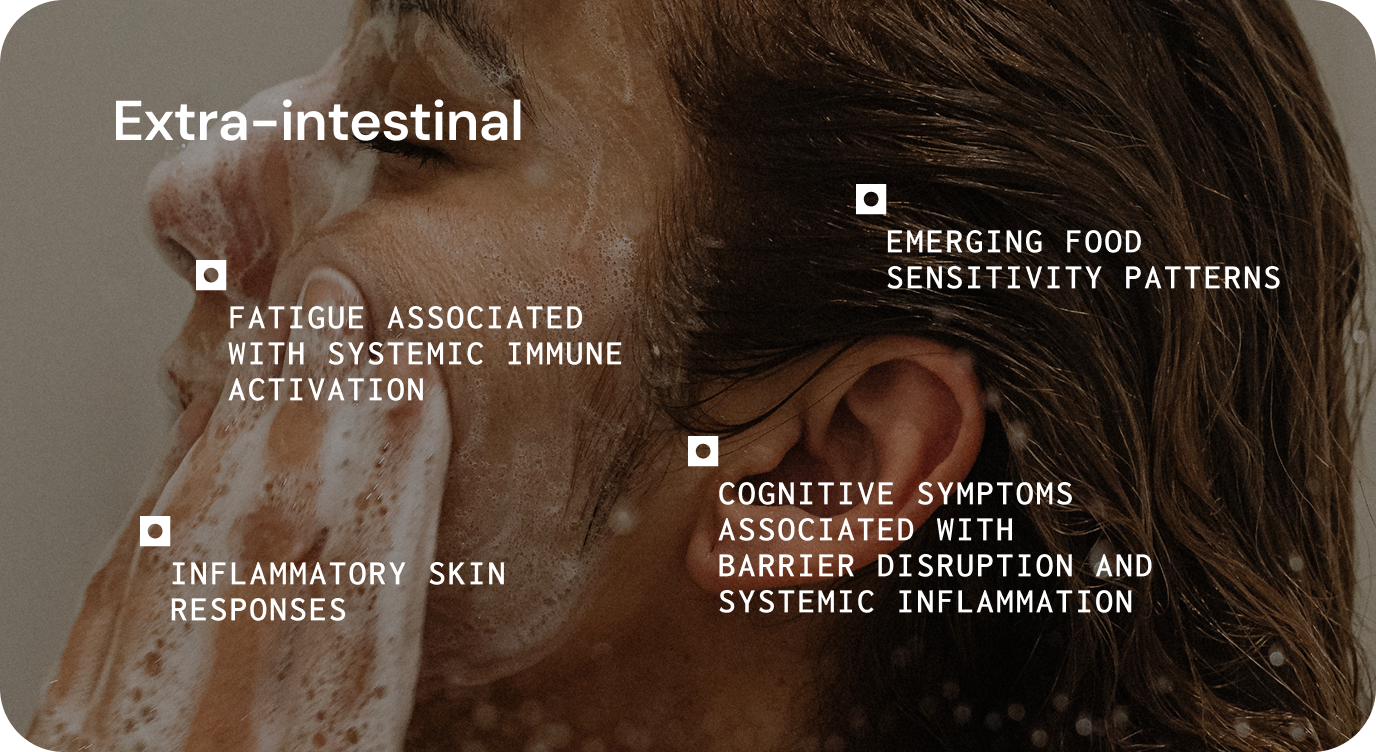
The gut microbiome can influence chronic disease by affecting core functions such as metabolism, immune regulation, gut barrier integrity, and systemic signalling.1 This helps explain why microbial imbalance may contribute to both gastrointestinal symptoms and wider systemic presentations. 1,6,7,8
In practice, these disturbances may present as persistent inflammatory symptoms, heightened sensitivity to dietary triggers, altered bowel habits, fatigue, or variable response to established treatments.1,6,7
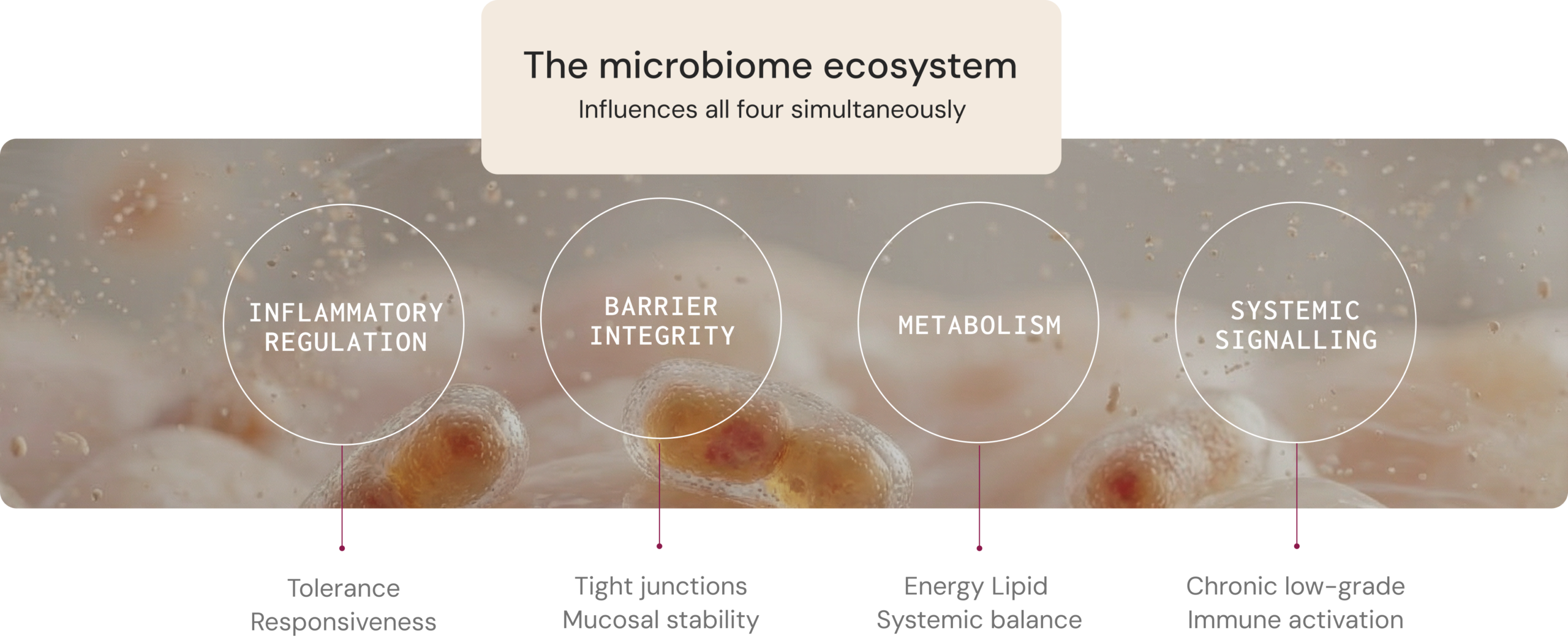
Four core physiological functions
influenced by the microbiome
The microbiome influences health through measurable host–microbe interactions that affect metabolism, immune regulation, gut barrier integrity, and systemic signalling. Understanding these mechanisms can help clinicians interpret how microbial ecology may contribute to symptom patterns and disease progression.1
Immune
regulation
Microbial components and metabolites interact continuously with immune cells within the intestinal mucosa. Short-chain fatty acids and other microbial metabolites influence regulatory T-cell activity, cytokine production and immune signalling pathways.6
IBD9, type 1 diabetes10, rheumatoid arthritis10
Gut barrier
function
Microbial metabolites support epithelial energy metabolism, tight junction stability, and mucosal signalling. When disrupted, increased permeability and altered mucosal signalling may contribute to immune activation and gastrointestinal symptoms.7,11
IBS12, IBD9, coeliac disease7
Metabolism
Microbial metabolites interact with host receptors involved in glucose and appetite regulation, lipid metabolism and inflammatory signalling pathways. These compounds influence endocrine signalling, energy metabolism and inflammation.8
Obesity, type 2 diabetes, NAFLD1
Systemic
signalling
Microbial metabolites interact with host receptors and influence signalling pathways beyond the gut, including enteroendocrine signalling, neural circuits, and immune pathways.11
Cardiovascular disease, Parkinson’s disease, type 2 diabetes13,14
Simplified microbiome models are not sufficient for effective care
Wellness framing
“Good vs bad bacteria.” Rebalance with a single product.
One-size-fits-all interventions. No consideration of individual ecology or patient context.
Clinical science
Dynamic ecosystems. Functional capacity and ecological structure. Strain-level differences. Context-dependent metabolic effects. Bidirectional microbiome signalling.
Two individuals taking the same probiotic or dietary intervention may experience different outcomes depending on their existing microbial ecology, metabolic pathways and host physiology.15 Without understanding this context, interventions may fail to address the mechanisms contributing to symptoms or disease progression.
A clinically responsible framework recognises individual variation, strain-level functional differences, context-dependent metabolic effects and bidirectional host–microbe signalling.2,3,15,16
Key Takeaway
The microbiome is directly involved in physiological functions central to chronic disease, including metabolism, immune regulation, gut barrier integrity, and systemic signalling.1 It is a biologically active system embedded within human physiology.
Understanding microbial ecosystem function is therefore an important component of modern clinical medicine and interpreting how the microbiome influences human health.17

- Fan Y, Pedersen O. Gut microbiota in human metabolic health and disease. Nat Rev Microbiol. 2021;19(1):55–71. https://doi.org/10.1038/s41579-020-0433-9
- Costello EK, Stagaman K, Dethlefsen L, Bohannan BJ, Relman DA. The application of ecological theory toward an understanding of the human microbiome. Science. 2012;336(6086):1255–62.
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–14.
- Dethlefsen L, Relman DA. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc Natl Acad Sci USA. 2011;108(Suppl 1):4554–61.
- Zhou X, Shen X, Johnson JS, et al. Longitudinal profiling of the microbiome at four body sites reveals core stability and individualized dynamics during health and disease. Cell Host Microbe. 2024;32(4):506–526.e9.
- Tan J, McKenzie C, Potamitis M, Thorburn AN, Mackay CR, Macia L. The role of short-chain fatty acids in health and disease. Adv Immunol. 2014;121:91–119.
- Ornelas A, Dowdell AS, Lee JS, Colgan SP. Microbial Metabolite Regulation of Epithelial Cell-Cell Interactions and Barrier Function. Cells. 2022;11(6):944.
- Koh A, De Vadder F, Kovatcheva-Datchary P, Bäckhed F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell. 2016;165(6):1332–1345. https://doi.org/10.1016/j.cell.2016.05.041
- Lloyd-Price J, Arze C, Ananthakrishnan AN et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature. 2019;569(7758):655–662. https://doi.org/10.1038/s41586-019-1237-9
- Honda K, Littman DR. The microbiota in adaptive immune homeostasis and disease. Nature. 2016;535(7610):75–84. https://doi.org/10.1038/nature18848
- Carabotti M, Scirocco A, Maselli MA, Severi C. The gut-brain axis: interactions between enteric microbiota, central and enteric nervous systems. Ann Gastroenterol. 2015;28(2):203–209.
- Staudacher HM, Mikocka-Walus A, Ford AC. Common mental disorders in irritable bowel syndrome: pathophysiology, management, and considerations for future randomised controlled trials. Lancet Gastroenterol Hepatol. 2021;6(5):401–410. https://doi.org/10.1016/S2468-1253(20)30363-0
- de Vos WM, Tilg H, Van Hul M, Cani PD. Gut microbiome and health: mechanistic insights. Gut. 2022;71(5):1020–1032. https://doi.org/10.1136/gutjnl-2021-326789
- Cryan JF, O’Riordan KJ, Cowan CSM, et al. The microbiota-gut-brain axis. Physiol Rev. 2019;99(4):1877–2013. https://doi.org/10.1152/physrev.00018.2018
- Leshem A, Segal E, Elinav E. The Gut Microbiome and Individual-Specific Responses to Diet. mSystems. 2020;5(5):e00665-20.
- Zmora N, Suez J, Elinav E. You are what you eat: diet, health and the gut microbiota. Nat Rev Gastroenterol Hepatol. 2019;16(1):35–56. https://doi.org/10.1038/s41575-018-0061-2
- Lynch SV, Pedersen O. The human intestinal microbiome in health and disease. N Engl J Med. 2016;375(24):2369–2379. https://doi.org/10.1056/NEJMra1600266
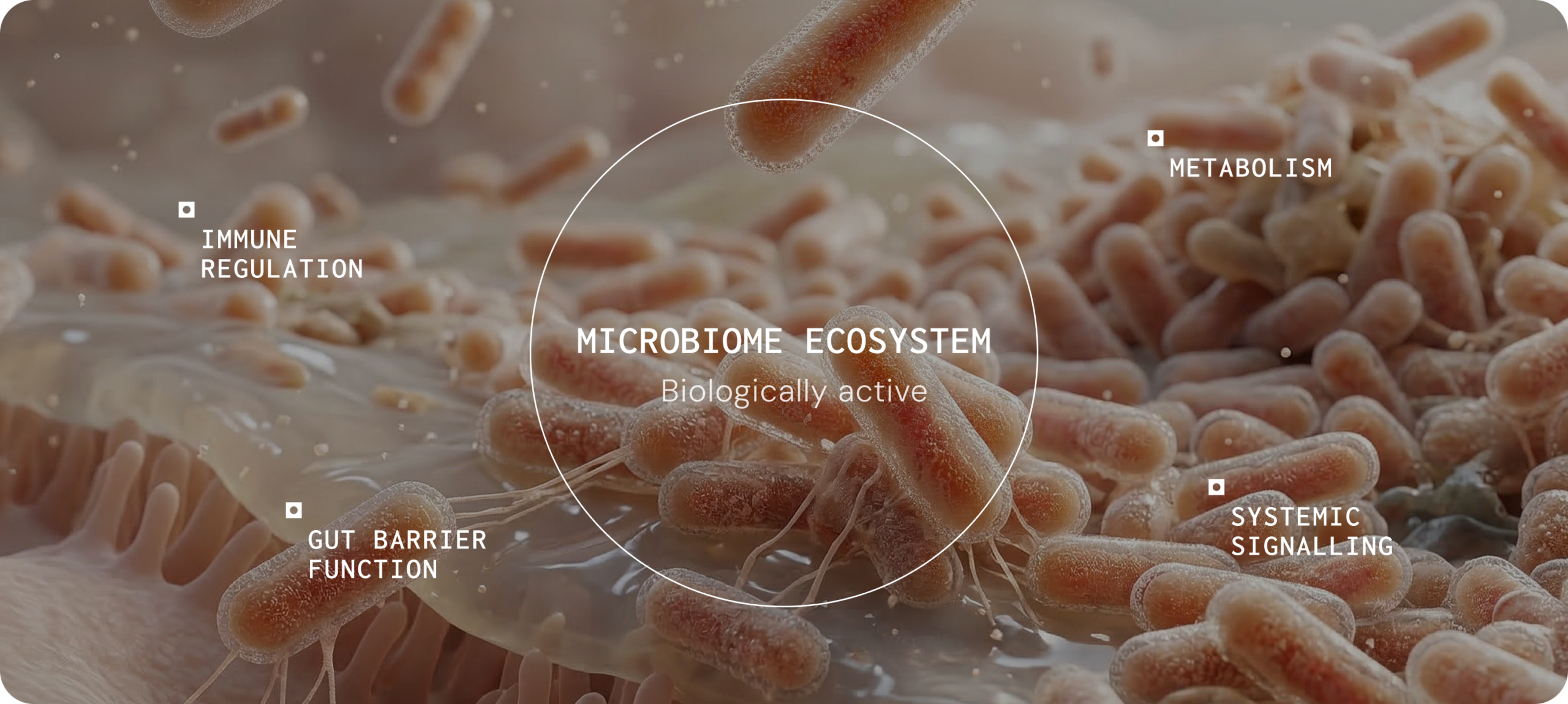
When microbial activity influences these systems, it can contribute to symptom persistence, inflammatory burden and variability in treatment response. In practice, clinicians may therefore consider whether host–microbe interactions are influencing a patient’s presentation, particularly in complex or chronic conditions where symptoms cannot be fully explained by structural pathology alone.
Integrated across core disciplines
Immunology
Immune education, inflammatory regulation, mucosal immunity
Gastroenterology
Barrier integrity, mucosal immune responses, motility
Endocrinology
Hormone regulation, appetite signalling, glucose metabolism
Metabolic medicine
Energy metabolism, lipid metabolism, systemic inflammatory tone
Positioning the microbiome within clinical medicine also requires adherence to the evidentiary and regulatory standards that govern medical practice. Clinical claims must be supported by reproducible evidence and remain proportionate to what current data can demonstrate. This is why microbiome research in medical contexts avoids overstating conclusions and remains grounded in established biological mechanisms.
Recognising the microbiome as clinically relevant does not mean attributing disease to microbial imbalance alone. Rather, it represents an additional physiological layer that interacts with systems already central to clinical medicine.
This is where microbiome science becomes clinically relevant. When microbial activity influences these systems, it can contribute to symptom persistence, inflammatory burden and variability in treatment response.
In practice, clinicians may therefore consider whether host–microbe interactions are influencing a patient’s presentation, particularly in complex or chronic conditions where symptoms cannot be fully explained by structural pathology alone.
Microbiome research is no longer about detection, it’s about function
Over the past two decades, microbiome research has moved from simply identifying which microbes are present to understanding what they do and how they interact with the body. Large population studies have shown that while microbial species differ widely between individuals, many core metabolic functions are preserved.2,3
EARLY 2000S
Species cataloguing and taxonomic classification
2010’s
Functional profiling and metagenomic sequencing
current
Clinical integration and longitudinal intervention studies
Early Microbiome Research
- Focused on identifying which microbial species are present
- Relied on organism detection and taxonomic classification
- Highlighted large species differences between individuals
- Limited ability to assess microbial function
- Mostly cross-sectional studies describing composition
Current Microbiome Research
- Focuses on what microbes do and how they interact with physiology
- Examines microbial metabolic activity and functional pathways
- Shows core metabolic functions are conserved across populations
- Metagenomic sequencing enables functional profiling
- Longitudinal studies examine response to diet and medication
Core physiological functions influenced by the microbiome
The microbiome influences health through measurable host–microbe interactions that affect metabolism, immune regulation, gut barrier integrity, and systemic signalling. Understanding these mechanisms can help clinicians interpret how microbial ecology may contribute to symptom patterns and disease progression.1
Microbial components and metabolites interact continuously with immune cells within the intestinal mucosa, including gut-associated lymphoid tissue. Short-chain fatty acids and other microbial metabolites influence regulatory T-cell activity, cytokine production and immune signalling pathways.8
Microbial signals support immune tolerance to dietary antigens and commensal microbes while preventing excessive inflammatory activation.
Persistent inflammatory symptoms, heightened immune sensitivity to dietary triggers, mucosal inflammation, exacerbation of immune-mediated gastrointestinal disorders.
Associated conditions
BD14, type 1 diabetes10, rheumatoid arthritis10
The intestinal epithelium is metabolically active and responsive to microbial signals. Microbial metabolites support epithelial energy metabolism, tight junction stability, and mucosal signalling, helping maintain a stable interface between luminal microbes and host tissues. When this regulation is disrupted, increased permeability and altered mucosal signalling may contribute to immune activation and gastrointestinal symptoms.9
The gut barrier functions as a stable interface between luminal microbes and host tissues, allowing efficient nutrient absorption while limiting inappropriate immune activation.
Increased intestinal permeability, impaired epithelial integrity, and altered mucosal signalling, which may contribute to inappropriate immune activation and gastrointestinal symptoms.
Associated conditions
IBS3, IBD10, coeliac disease10
Microbial metabolites interact with host receptors and influence signalling beyond the gut, including enteroendocrine signalling, neural circuits, and immune pathways. The gut and brain communicate through nerves, hormones, and immune signals. Gut microbes can influence these pathways by producing or modifying neuroactive compounds and by modulating inflammation levels.11,12
Coordinated neuroimmune and enteroendocrine signalling, stable gut–brain communication, balanced stress response.
Altered stress response, neuroinflammation, disrupted gut–brain communication, variable systemic signalling.
Associated conditions
Cardiovascular disease, type 2 diabetes, Parkinson’s disease6,10
Microbial metabolites interact with host receptors involved in glucose and appetite regulation, lipid metabolism and inflammatory signalling pathways. Short-chain fatty acids and related compounds influence endocrine signalling, energy metabolism and inflammation.12
Stable energy metabolism, balanced inflammatory responses, appropriate appetite and glucose regulation.
Metabolic dysregulation, fatigue associated with inflammatory activity, altered appetite regulation, variable response to dietary or metabolic interventions.
Associated conditions
Obesity, type 2 diabetes, NAFLD3
Physiological Function Summary
Immune regulation
Barrier integrity
Metabolic signalling
Systemic signalling
The microbiome influences chronic disease through core physiological functions
Chronic diseases rarely arise from disruption of a single biological pathway. Instead, they typically involve dysregulation across interconnected physiological functions, including metabolism, immune regulation, gut barrier integrity, and systemic signalling.1
In clinical practice, dysbiosis should not be interpreted simply as the presence of a pathogen or depletion of a single “beneficial” organism. More commonly, dysbiosis reflects broader shifts in microbial ecology and metabolic output that interact with existing clinical vulnerabilities.

What this looks like in practice
inflammatory regultion
Microbial metabolites, including short-chain fatty acids (SCFAs) and some indole derivatives, play an important role in immune regulation. These compounds influence immune cell differentiation, cytokine production and inflammatory signalling.8 Reduced SCFA-producing capacity has been associated with conditions characterised by immune dysregulation, including inflammatory bowel disease, autoimmune disorders and the low-grade systemic inflammation observed in metabolic syndrome.5,8
Clinical presentations
The following clinical presentations have been associated with microbiome-related immune dysregulation. Causal relationships are not established for all; these associations should be interpreted in the context of each patient’s clinical picture.1,8
Barrier integrity
The intestinal epithelium forms a critical interface between luminal microbial communities and the host immune system. Microbial metabolites such as butyrate support epithelial energy metabolism, tight junction integrity and mucosal immune signalling.9 When epithelial barrier regulation is disrupted, microbial products such as hexa-acylated lipopolysaccharides (hexa-LPS) may cross the intestinal barrier more readily, promoting immune activation and inflammatory signalling.1,9
Clinical presentations
The following clinical presentations have been associated with microbial disruption in the literature.
Causal relationships are not established for all; these associations should be interpreted in the context of each patient’s clinical picture
Metabolism
Gut microbes participate in nutrient metabolism and energy regulation through the production of bioactive metabolites that interact with metabolic pathways. Alterations in microbial metabolic activity have been associated with differences in glycaemic control, energy homeostasis and systemic inflammatory tone.12
Clinical presentations 1,13
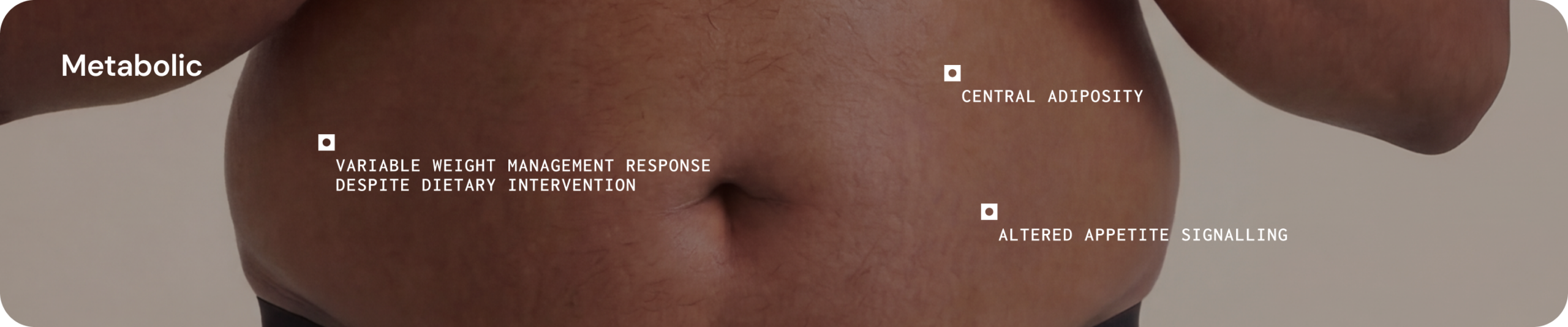
Systemic signalling
Microbial products can influence signalling beyond the gut through several recognised routes, including enteroendocrine signalling, gut-innervating afferent nerves such as the vagus nerve, and immune pathways.1,13 Short-chain fatty acids, tryptophan-derived metabolites, and hexa-acylated lipopolysaccharides can act on enteroendocrine and immune cells, modulate cytokine signalling, and influence neuronal communication between the gut and brain.13
These interactions are bidirectional. Factors such as diet, immune status, medications and environmental exposures continuously shape microbial ecology, while microbial metabolites in turn influence immune regulation, gut barrier integrity, metabolism and systemic signalling. Patients with similar diagnoses can respond differently to the same interventions — differences in microbial metabolism are increasingly recognised as a contributing factor.1,8,9
Did you know?
These interactions are bidirectional. Diet, immune status, medication exposure and environmental factors continuously shape microbial ecology, while microbial metabolites in turn influence immune function, epithelial integrity and metabolic regulation. Patients with similar diagnoses can respond differently to the same interventions — differences in microbial metabolism are increasingly recognised as a contributing factor.8,9,10
Simplified microbiome models are not sufficient for clinical practice
Wellness messaging often reduces the microbiome to simplified concepts such as “good versus bad bacteria” or the idea that gut health can be restored by simply “rebalancing” microbial populations. These narratives are appealing because they are easy to communicate, but they do not reflect how microbial ecosystems function within human physiology.
Clinical science approaches the microbiome differently. Rather than focusing on individual organisms classified as beneficial or harmful, it considers microbial communities as dynamic ecosystems whose effects depend on ecological structure, functional capacity and host context.3
| Wellness framing
“Good vs bad bacteria.” Rebalance with a single product. Clinical science Dynamic ecosystems. Functional capacity and ecological structure. Strain-level differences. Context-dependent metabolic effects. Bidirectional microbiome signalling |
This approach also reflects the clinical and regulatory standards that govern medical practice. In medical contexts, health claims must be supported by robust scientific evidence, which is why clinical microbiome research avoids overstating conclusions and remains grounded in established biological mechanisms.
As a result, interventions based purely on simplified microbial classifications may produce inconsistent outcomes. Two individuals taking the same probiotic, dietary intervention or supplement may experience different effects depending on their existing microbial ecology, metabolic pathways and host physiology.16
When microbiome interventions are selected without consideration of the clinical presentation, they may not target the biological mechanisms contributing to a patient’s symptoms. In some cases, this may result in minimal benefit, transient improvement or variability in response between individuals.
A clinically responsible framework therefore recognises several principles supported by microbiome research:
- Substantial variation: there is substantial variation in microbial composition between individuals, even among healthy population
- Functional differences at species and strain level that influence metabolic activity
- Context-dependent metabolic effects rather than outcomes determined by the presence of a single organism
- Bidirectional signalling between host physiology and microbial activity 3,4,16,17
Key Takeaway
The gut microbiome is a measurable, modifiable part of human physiology. It functions as an ecosystem — and like any ecosystem, its health depends on the balance, diversity, and functional capacity of the community as a whole, not just the presence or absence of individual organisms.
For clinicians, the most useful insight from the microbiome is often functional rather than taxonomic: what the microbial community is producing, how that interacts with the patient’s immune system, gut barrier, and metabolic regulation, and how those outputs may be shifting in the context of that individual’s diet, medications, and clinical picture.
For clinicians, microbiome insights provide an additional lens for understanding systemic regulation, particularly in chronic and complex symptom patterns.
As our understanding of the microbiome deepens, so does the opportunity to use it as a meaningful part of clinical assessment, particularly for patients with chronic or multi-system presentations where traditional markers tell only part of the story.

1. Fan Y, Pedersen O. Gut microbiota in human metabolic health and disease. Nat Rev Microbiol. 2021;19(1):55–71. https://doi.org/10.1038/s41579-020-0433-9
2. Lynch SV, Pedersen O. The human intestinal microbiome in health and disease. N Engl J Med. 2016;375(24):2369–2379. https://doi.org/10.1056/NEJMra1600266
3. Wilde J, Slack E, Foster KR. Host control of the microbiome: Mechanisms, evolution, and disease. Science. 2024;385(6706):eadi3338. https://doi.org/10.1126/science.adi3338
4. Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–14.
5. Mann ER, Lam YK, Uhlig HH. Short-chain fatty acids: linking diet, the microbiome and immunity. Nat Rev Immunol. 2024;24(8):577–595. https://doi.org/10.1038/s41577-024-01014-8
6. de Vos WM, Tilg H, Van Hul M, Cani PD. Gut microbiome and health: mechanistic insights. Gut. 2022;71(5):1020–1032. https://doi.org/10.1136/gutjnl-2021-326789
7. Zhou X, Shen X, Johnson JS, et al. Longitudinal profiling of the microbiome at four body sites reveals core stability and individualized dynamics during health and disease. Cell Host Microbe. 2024;32(4):506–526.e9.
8. Hays KE, Pfaffinger JM, Ryznar R. The interplay between gut microbiota, short-chain fatty acids, and implications for host health and disease. Gut Microbes. 2024;16(1):2393270. https://doi.org/10.1080/19490976.2024.2393270
9. Ornelas A, Dowdell AS, Lee JS, Colgan SP. Microbial Metabolite Regulation of Epithelial Cell-Cell Interactions and Barrier Function. Cells. 2022;11(6):944.
10. Honda K, Littman DR. The microbiota in adaptive immune homeostasis and disease. Nature. 2016;535(7610):75–84. https://doi.org/10.1038/nature18848
11. Koh A, De Vadder F, Kovatcheva-Datchary P, Bäckhed F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell. 2016;165(6):1332–1345. https://doi.org/10.1016/j.cell.2016.05.041
12. Muramatsu MK, Winter SE. Nutrient acquisition strategies by gut microbes. Cell Host Microbe. 2024;32(6):863–874. https://doi.org/10.1016/j.chom.2024.05.011
13. Cryan JF, O’Riordan KJ, Cowan CSM, et al. The microbiota-gut-brain axis. Physiol Rev. 2019;99(4):1877–2013. https://doi.org/10.1152/physrev.00018.2018
14. Lloyd-Price J, Arze C, Ananthakrishnan AN et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature. 2019;569(7758):655–662. https://doi.org/10.1038/s41586-019-1237-9
15. Staudacher HM, Mikocka-Walus A, Ford AC. Common mental disorders in irritable bowel syndrome: pathophysiology, management, and considerations for future randomised controlled trials. Lancet Gastroenterol Hepatol. 2021;6(5):401–410. https://doi.org/10.1016/S2468-1253(20)30363-0
16. Leshem A, Segal E, Elinav E. The Gut Microbiome and Individual-Specific Responses to Diet. mSystems. 2020;5(5):e00665-20.
17. Zmora N, Suez J, Elinav E. You are what you eat: diet, health and the gut microbiota. Nat Rev Gastroenterol Hepatol. 2019;16(1):35–56. https://doi.org/10.1038/s41575-018-0061-2
Microbial diversity protects against pathogens through colonisation resistance
One of the microbiome’s key roles is protecting against harmful microorganisms, a process called colonisation resistance. In a diverse, well-balanced community, many species compete for space and nutrients, leaving few open niches for potential pathogens to occupy. Other mechanisms of colonisation resistance include the production of acidic metabolites that lower luminal pH, antimicrobial compounds, and metabolites that support barrier function and host immunity.4,5,6,7
Did you know?
When diversity is reduced — after antibiotics, severe illness or restrictive diets — this protective network can weaken. With fewer competitors and less functional redundancy (fewer microbes able to perform the same protective functions), opportunistic organisms may gain a foothold and, in some cases, establish infection or persistent colonisation.4,7
Clinical example
Impaired colonisation resistance has been linked to greater susceptibility to post-infectious IBS following gastroenteritis, with some individuals experiencing long-term symptoms after an acute infection. 8,9

Diet determines microbial outputs
The same microbes can produce different metabolites
What microbes do is strongly influenced by what they are fed. Many dietary fibres cannot be digested by human enzymes but can be fermented by gut microbes, producing health-promoting short-chain fatty acids (SCFAs). 10,11 Microbes also ferment undigested protein and amino acids in the distal colon, producing metabolites that may be harmful or beneficial. 10

Critically, the gut microbiota is metabolically flexible. Microbes that produce beneficial metabolites from dietary fibre can shift towards less favourable outputs when the diet is low in fermentable carbohydrates and high in protein or fat. 10,13
The type and diversity of fibre matter as much as total fibre.
Microbial metabolites shape immune
function across the lifespan
From birth, the immune system learns to distinguish between threats and harmless exposures. Early contact with diverse microbes supports the development of regulatory pathways that reduce the risk of over-reactive responses. Disruptions in early-life microbial exposure have been associated with increased risk of allergies, asthma, inflammatory bowel disease and autoimmune conditions.21
In adulthood, the microbiome continues to shape immune function: Microbiota-derived SCFAs can promote regulatory immune responses. 14,24 Similarly, tryptophan-derived metabolites can modulate innate immune responses. 25 Lipopolysaccharides (bacterial cell wall components) can either trigger or dampen inflammatory signalling. 26 When the microbiome is balanced, these signals usually support a steady, controlled immune state. 27
Clinical example
In oncology, gut microbiome composition has been linked to better or worse responses to immunotherapy, suggesting microbial immune modulation can influence treatment outcomes.28
Health outcomes depend on community cooperation, not individual organisms
Many clinically relevant metabolites are not produced by a single microbe acting alone. Instead, they arise through cross-feeding, where one species’ metabolic by-products become another species’ fuel. People with very different species profiles can still display similar functional capacities: different microbial “teams” can lead to similar metabolic outcomes.3,29
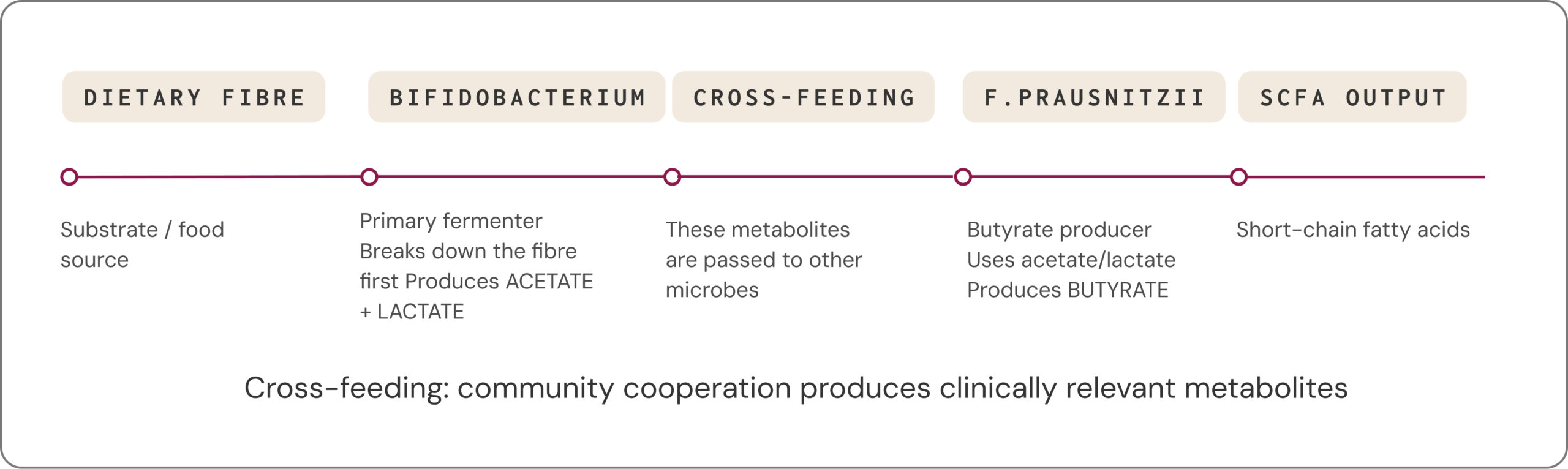
Clinical example
After antibiotic courses, some individuals develop prolonged bloating, pain and altered bowel habits. Studies have reported disrupted cross-feeding networks and lower SCFA production in these settings, consistent with loss of key metabolic partners and reduced ecosystem resilience.30

Functional potential only becomes functional reality when the right conditions align
Modern sequencing technologies can estimate the microbiome’s functional potential by identifying genes and pathways — a valuable starting point. However, this potential must be realised through the right conditions, and especially fuel source availability.3
Because healthy people can have very different species profiles, it is difficult to define a single “ideal” microbiome based only on who is present.
It is more practical to think in terms of functional balance: a state where activities that support health are robust, while pathways that generate potentially harmful metabolites are kept in check.31
FUNCTIONAL POTENTIAL
Genes and pathways are present
RIGHT CONDITIONS
Available fibre, substrates, microbial cooperation, gut environment
FUNCTIONAL REALITY
Metabolites are actively produced
Key Takeaway
The microbiome is a dynamic ecosystem whose functions — not just its composition — shape human health. Diverse communities support colonisation resistance, diet along with other factors determine metabolic outputs, and health outcomes depend on community cooperation and functional balance rather than individual species.
Functional potential only becomes functional reality when the right fuels and conditions align. A functional, ecosystem-based lens supports more personalised, targeted strategies to maintain or restore health.

- Coyte KZ, Schluter J, Foster KR. The ecology of the microbiome: networks, competition, and stability. Science. 2015;350:663–666.
- Schroeder BO, Bäckhed F. Signals from the gut microbiota to distant organs in physiology and disease. Nat Med. 2016;22:1079–1089.
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–214.
- Sorbara MT, Pamer EG. Interbacterial mechanisms of colonization resistance. Mucosal Immunol. 2019;12:1–9.
- Caballero-Flores G, Pickard JM, Núñez G. Microbiota-mediated colonization resistance. Nat Rev Microbiol. 2023;21:347–360.
- Kamada N, Chen GY, Inohara N, Núñez G. Control of pathogens and pathobionts by the gut microbiota. Nat Immunol. 2013;14:685–690.
- Buffie CG, Pamer EG. Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol. 2013;13:790–801.
- Jalanka J, Gunn D, Singh G, Krishnasamy S, Lingaya M, Crispie F, et al. Postinfective bowel dysfunction following Campylobacter enteritis. Gut. 2023;72:451–459.
- Marshall JK, Thabane M, Garg AX, Clark WF, Moayyedi P, Collins SM. Eight year prognosis of postinfectious IBS. Gut. 2010;59:605–611.
- Rowland I, Gibson G, Heinken A, Scott K, Swann J, Thiele I, et al. Gut microbiota functions: metabolism of nutrients and other food components. Eur J Nutr. 2018;57:1–24.
- Koh A, De Vadder F, Kovatcheva-Datchary P, Bäckhed F. From dietary fiber to host physiology: SCFAs as key bacterial metabolites. Cell. 2016;165:1332–1345.
- Peng L, Li ZR, Green RS, Holzman IR, Lin J. Butyrate enhances the intestinal barrier. J Nutr. 2009;139:1619–1625.
- Gilbert MS, Ijssennagger N, Kies AK, van Mil SWC. Protein fermentation in the gut. Am J Physiol Gastrointest Liver Physiol. 2018;315:G159–G170.
- Corrêa-Oliveira R, Fachi JL, Vieira A, Sato FT, Vinolo MAR. Regulation of immune cell function by short-chain fatty acids. Clin Transl Immunol. 2016;5:e73.
- Alexeev EE, Lanis JM, Kao DJ, Campbell EL, Kelly CJ, Battista KD, et al. Microbiota-derived indole metabolites promote intestinal homeostasis through regulation of interleukin-10 receptor. Am J Pathol. 2018;188:1183–1194.
- Canfora EE, Jocken JW, Blaak EE. Short-chain fatty acids in control of body weight and insulin sensitivity. Nat Rev Endocrinol. 2015;11:577–591.
- Sinha AK, Laursen MF, Brinck JE, Rybtke ML, Hjørne AP, Procházková N, et al. Dietary fibre directs microbial tryptophan metabolism via metabolic interactions in the gut microbiota. Nat Microbiol. 2024;9:1964–1978.
- Teichmann J, Cockburn DW. Butyrate production dependent on resistant starch source. Front Microbiol. 2021;12:640253.
- Venkataraman A, Sieber JR, Schmidt AW, Waldron C, Theis KR, Schmidt TM. Variable responses to dietary supplementation with resistant starch. Microbiome. 2016;4:33. 20. Ďásková N, Modos I, Krbcová M, Kuzma M, Pelantová H, Hradecký J, et al. Multi-omics signatures predict metabolic response to dietary inulin. Nutr Diabetes. 2023;13:7.
- Gensollen T, Iyer SS, Kasper DL, Blumberg RS. How colonization by microbiota in early life shapes the immune system. Science. 2016;352:539–544.
- Wernroth M-L, Fall K, Svennblad B, Ludvigsson JF, Sjölander A, Almqvist C, et al. Early childhood antibiotic treatment is associated with risk of type 1 diabetes. Diabetes Care. 2020;43:991–999.
- Kronman MP, Zaoutis TE, Haynes K, Feng R, Coffin SE. Antibiotic exposure and IBD development among children. Pediatrics. 2012;130:e794–e803.
- Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, deRoos P, et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature. 2013;504:451–455.
- Zelante T, Iannitti RG, Cunha C, De Luca A, Giovannini G, Pieraccini G, et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor. Immunity. 2013;39:372–385.
- Mohr AE, Crawford M, Jasbi P, Fessler S, Sweazea KL. Lipopolysaccharide and the gut microbiota: considering structural variation. FEBS Lett. 2022;596:849–875.
- Zheng D, Liwinski T, Elinav E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020;30:492–506.
- Baruch EN, Youngster I, Ben-Betzalel G, Ortenberg R, Lahat A, Katz L, et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma. Science. 2021;371:602–609.
- Culp EJ, Goodman AL. Cross-feeding in the gut microbiome. Cell Host Microbe. 2023;31:485–499.
- Palleja A, Mikkelsen KH, Forslund SK, Kashani A, Allin KH, Nielsen T, et al. Recovery of gut microbiota following antibiotic exposure. Nat Microbiol. 2018;3:1255–1265.
- Tiffany CR, Bäumler AJ. Dysbiosis: from fiction to function. Am J Physiol Gastrointest Liver Physiol. 2019;317:G602–G608.