The microbiome isn’t a list of species.
It’s a living ecosystem. It should not be measured by testing fragments in isolation. The whole picture matters.
A healthy gut microbiome requires a balanced ecosystem
Like any ecosystem, a healthy gut microbiome is a balanced one. Evaluating that balance cannot be reduced to a list of well-known species. It emerges from complex microbial interactions across the whole ecosystem, from diversity to environment

Diversity

Relative abundance

Functional capacity

Physiological environment
Most tests only measure a partial view of that ecosystem
Most testing technologies measure only fragments of this ecosystem, evaluating a partial list of well-known species and attempt to infer "balance" from that partial view. If the clinical question is ecosystem balance, the measurement must reflect the ecosystem.
Species-level detection
Measures the presence or absence of individual organisms in isolation - no diversity data, no functional insight, no understanding of how the community behaves as a whole.
Whole ecosystem assessment
Captures the full community picture: diversity, relative abundance, functional capacity, and the physiological conditions that shape how the microbiome actually functions.
Looking for the deep dive?
Read the full article: Measuring the microbiome as an ecosystem
Partial testing can’t reliability answer
critical clinical questions
How does this organism’s abundance compare to the overall ecosystem?
Is the ecosystem resilient or vulnerable to instability? ²³
Is functional capacity reduced?
Is it contributing to an anti-inflammatory or pro-inflammatory state?

Ecosystem-level assessment unlocks critical patient insights.
In practice, microbiome testing is about more than detecting which species are present. The goal is interpretation: understanding how the microbiome may be influencing symptoms, inflammation, gut barrier function, and metabolic signalling

Chronic symptoms

Inflammatory resistance
Gut barrier integrity

Metabolic signalling
Testing technologies vary in what they can measure and reveal
Culture
Useful for pathogen detection - not ecosystem assessment.
- Culture can identify certain infectious organisms.
- Identifies organisms that grow in lab conditions
- Misses many microbes that cannot be cultured
- Provides no insight into diversity or function¹
Targeted PCR / qPCR
Highly sensitive, but only for selected targets.
- Excellent at confirming specific organisms
- Cannot assess broader community balance
- Does not detect organisms outside the selected panel.²
16S rRNA gene sequencing
Broader view — but limited resolution and no direct functional insight.
- Surveys bacterial communities without culture
- Often limited to genus-level resolution
- Does not directly measure functional genes or pathways⁴
Shotgun Metagenomics
Whole-community, species-level, function-aware measurement.
- Sequences all DNA in the sample
- Detects bacteria, fungi, archaea and other microbes
- Provides species-level identification
- Enables functional gene and pathway profiling⁵
Looking for the deep dive?
Read the full article: Measuring the microbiome as an ecosystem
The critical shift
Metagenomics enables ecosystem-level assessment. Because it captures the full genetic content of the community. ⁵ Not just selected fragments
Ecosystem-level assessment reveals functional capacity
Clinical impact is determined not just by which organisms are present, but by what the ecosystem is functionally doing within the body. We call this functional capacity.
Examples include
- Short-chain fatty acid production (e.g. butyrate), which supports immune regulation and barrier integrity.⁵
- Lipopolysaccharide (LPS) signalling potential from Gram-negative organisms, associated with inflammatory activation.⁸
- Mechanisms influencing gut barrier permeability.

Without insights into the functional capacity of the microbiome, clinicians are left interpreting microbial composition alone, which often fails to explain chronic symptoms or relapsing presentation.
Whole-ecosystem testing supports more informed clinical decision-making
Whole-ecosystem testing fundamentally changes how clinicians think about patient symptoms. It moves the clinical question from “what organism is present?” to “how is this ecosystem functioning, and what is it driving?” This is where clinical-grade testing comes in.

Clinical-grade testing requires more than technology
Whole-community sequencing is a necessary foundation, but clinical application requires more.
Metagenomics is only clinically meaningful when results are accurate, reproducible, and interpretable. For that, the technology has to be embedded in validated, accredited systems. Technology alone doesn't deliver clarity. Integrated systems do.
Validated sample collection and preservation
Accredited laboratory standards (e.g. ISO 15189)
High-quality bioinformatics pipelines⁷
Evidence-based interpretation frameworks
The most accurate picture requires the right combination of technologies
No single method answers every clinical question.
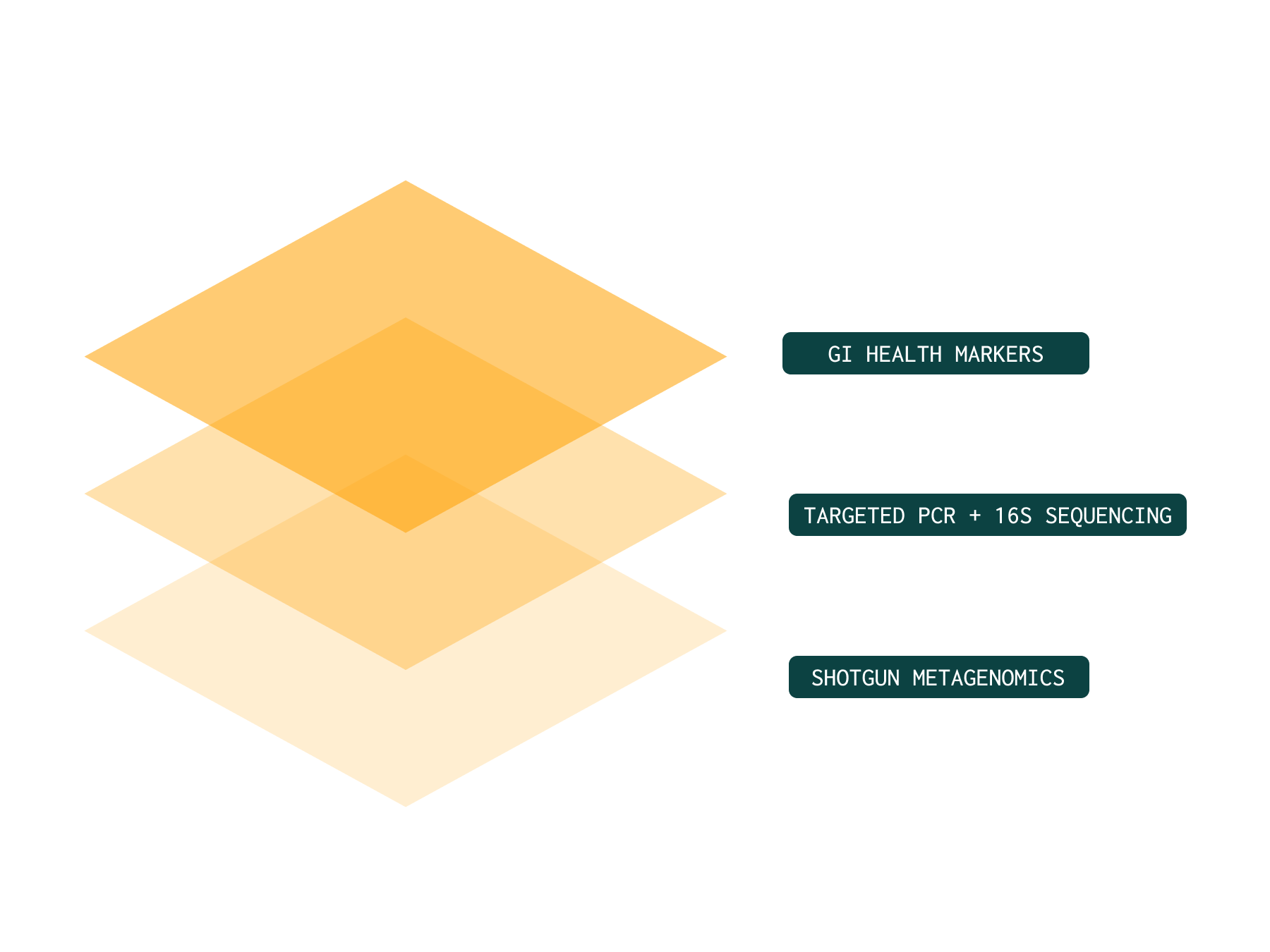
To deliver true ecosystem insight, testing must integrate:
Shotgun metagenomics for full community and functional capacity⁵
Validated Gut health markers to assess gut barrier function and inflammation
PCR for targeted pathogen confirmation
Practical summary
Microbiome balance is an ecosystem property within the gut environment. ¹²
- Partial testing measures fragments, not systems.
- Functional capacity determines clinical impact. ⁵
- Ecosystem-level understanding requires whole-community measurement.
- Clinical-grade clarity requires validated systems, not just sequencing.
- If the goal is confident, actionable insights, then the whole ecosystem must be
Costello EK et al. The application of ecological theory toward an understanding of the human microbiome. Science. 2012.
Coyte KZ et al. The ecology of the microbiome: networks, competition, and stability. Science. 2015.
Lozupone CA et al. Diversity, stability and resilience of the human gut microbiota. Nature. 2012.
Jovel J et al. Characterization of the gut microbiome using 16S or shotgun metagenomics. Front Microbiol. 2016.
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012.
Ranjan R et al. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. BBRC. 2016.
Quince C et al. Shotgun metagenomics, from sampling to analysis. Nat Biotechnol. 2017.
Zhernakova A et al. Population based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science. 2016.