The microbiome is a dynamic ecosystem, not a collection of individual species
Every person harbours a vast community of microorganisms — mostly bacteria, along with fungi, viruses and archaea — collectively known as the microbiome. This community is densest in the gut, especially the colon, where trillions of microbes interact with each other and with their human host. Rather than acting as isolated species, these microbes form an ecosystem that adjusts to diet, medications, infections, stress and other exposures.1,2
The functional outputs of this ecosystem depends on many moving parts, including: which microbes are present, how they interact, what fuels they receive, the gut environment, and which metabolic activities are turned on at any given time.1,3 In turn, these metabolic outputs shape host physiology and can promote health or, if out of balance, may contribute to symptoms and disease processes. 2
Microbial diversity protects against pathogens through colonisation resistance
One of the most important ways the microbiome supports health is by helping protect against harmful microorganisms, a process called colonisation resistance. In a diverse, well-balanced community, many species compete for space and nutrients, leaving few open niches for potential pathogens to occupy.
Production of acidic metabolites such as lactate and SCFAs lowers luminal pH, making the colon less hospitable for harmful bacteria. 6,7
Generation of bacteriocins and other compounds that directly inhibit competitors. 6
Production of metabolites that support the mucus layer and gut barrier, limiting pathogen and toxin access to the intestinal wall.
Promotion and priming of innate immune cells, enabling faster response to enteric infection.
Did you know?
When diversity is reduced — after antibiotics, severe illness or restrictive diets — this protective network can weaken. With fewer competitors and less functional redundancy (fewer microbes able to perform the same protective functions), opportunistic organisms may gain a foothold and, in some cases, establish infection or persistent colonisation.4,7
Clinical example
Impaired colonisation resistance has been linked to greater susceptibility to post-infectious IBS following gastroenteritis, with some individuals experiencing long-term symptoms after an acute infection. 8,9

Diet determines microbial outputs
The same microbes can produce different metabolites. What microbes do is strongly influenced by what they are fed. Many dietary fibres cannot be digested by human enzymes but can be fermented by gut microbes. 10 When bacteria ferment complex carbohydrates, they mainly produce health-promoting short-chain fatty acids such as acetate, propionate and butyrate. 11

Critically, the gut microbiota is metabolically flexible. Microbes that produce beneficial metabolites from dietary fibre can shift towards less favourable outputs when the diet is low in fermentable carbohydrates and high in protein or fat. Different fibres (resistant starch, inulin, arabinoxylans) favour different microbial groups and can shift SCFA profiles, so the type and diversity of fibre matter as much as total fibre intake.
Clinical example
As part of amino-acid fermentation, certain microbes can convert histidine into histamine. In some IBS cohorts, higher microbial histamine production has been associated with abdominal pain and visceral hypersensitivity. 21

Microbial metabolites shape immune function across the lifespan
Early-life training.
From birth, the immune system learns to distinguish between threats and harmless exposures. Microbes are key teachers
in this process. Early contact with a diverse set of microbes helps immune cells learn when to respond strongly and when to
remain tolerant, supporting the development of regulatory pathways that reduce the risk of over-reactive responses.22
Disruptions in early-life microbial exposure — frequent antibiotics, limited dietary diversity or lack of environmental microbial contact — have been associated with increased risk of allergies, asthma, inflammatory bowel disease and some autoimmune conditions later in life.23,24
BIRTH
Initial microbial colonisation begins immune education
EARLY CHILDHOOD
Diverse exposure builds tolerance and regulatory pathways
ADULTHOOD
Ongoing immune modulation through microbial metabolites and signalling
Ongoing immune modulation in adults
In adulthood, the microbiome continues to shape immune tone. Microbia products interact with pattern-recognition receptors and other immune sensors, sending signals that can either amplify or dampen inflammation.25
Promote regulatory T cells and reduce certain pro- inflammatory cytokines19,26
Signal through aryl hydrocarbon receptors influencing innate lymphoid cells and local immune responses27
Different forms of LPS bacterial cell wall components can either trigger pro-inflammatory signalling or elicit more regulatory immune responses. 28
When the microbiome is balanced, these signals usually support a steady, controlled immune state.29 When microbial composition and function are disrupted, signalling may become skewed in ways that can contribute to chronic inflammation in susceptible individuals.25,29
Clinical example
In oncology, gut microbiome composition has
been linked to better or worse responses to immunotherapy, suggesting microbial immune modulation can influence treatment outcomes.30
Health outcomes depend on community cooperation, not individual organisms
Cross-feeding and microbial cooperation.
Many clinically relevant metabolites are not produced by a single microbe acting alone. Instead, they arise through cross-feeding, where one species’ metabolic by-products become another species’ fuel. For example, fibre fermentation by Bifidobacterium spp. can generate acetate and lactate that is then converted into butyrate by butyrate-producing Firmicutes such as Faecalibacterium prausnitzii, Roseburia and Eubacterium.31
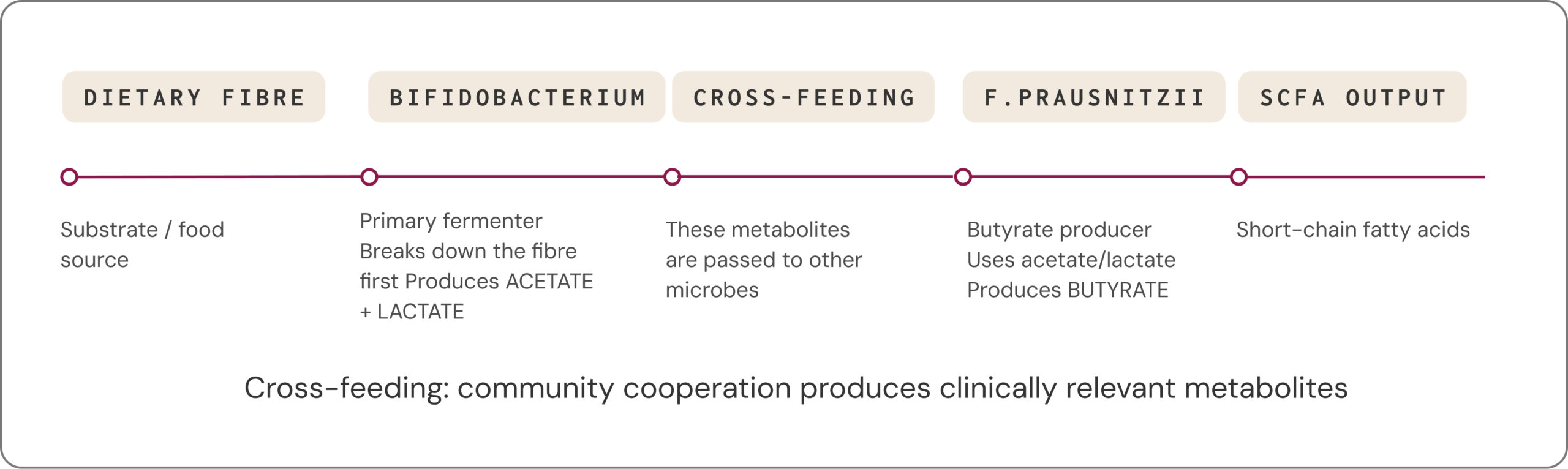
This distributed architecture also helps explain how microbial communities become more stable and resilient to disturbances, and how people with very different species profiles can still display similar functional capacities: different microbial “teams” can lead to similar metabolic outcomes.3,31
Competition and ecological balance
Alongside cooperation, microbes compete for resources and space. They may alter the local pH, secrete inhibitory molecules or more efficiently harvest available nutrients
These interactions influence which groups become dominant and which functions are emphasised. A community with a robust network of cooperative and competitive interactions tends to be more stable and resilient.1,6
After antibiotic courses, some individuals develop prolonged bloating, pain and altered bowel habits.
Studies have reported disrupted cross-feeding networks and lower SCFA production in these settings,
consistent with loss of key metabolic partners and reduced ecosystem resilience.29,30
Functional potential only becomes functional reality when the right conditions align
Modern sequencing technologies can estimate the microbiome’s functional potential by identifying genes and pathways — a valuable starting point. However, this potential must be realised through the right conditions, and especially fuel source availability.
What bacteria produce depends heavily on available substrates: when fermentable carbohydrates are plentiful, SCFA production often increases; but when dietary fibre is limited, proteolytic fermentation pathways can become more prominent.10
FUNCTIONAL POTENTIAL
Genes and pathways are present
RIGHT CONDITIONS
Available fibre, substrates, microbial cooperation, gut environment
FUNCTIONAL REALITY
Metabolites are actively produced
CLINICAL EXAMPLE
Individuals with higher butyrate-producing capacity may show greater response and metabolic benefit when diets provide matching fermentable fibres, illustrating how diet can “unlock” functional capacity.12,13,14

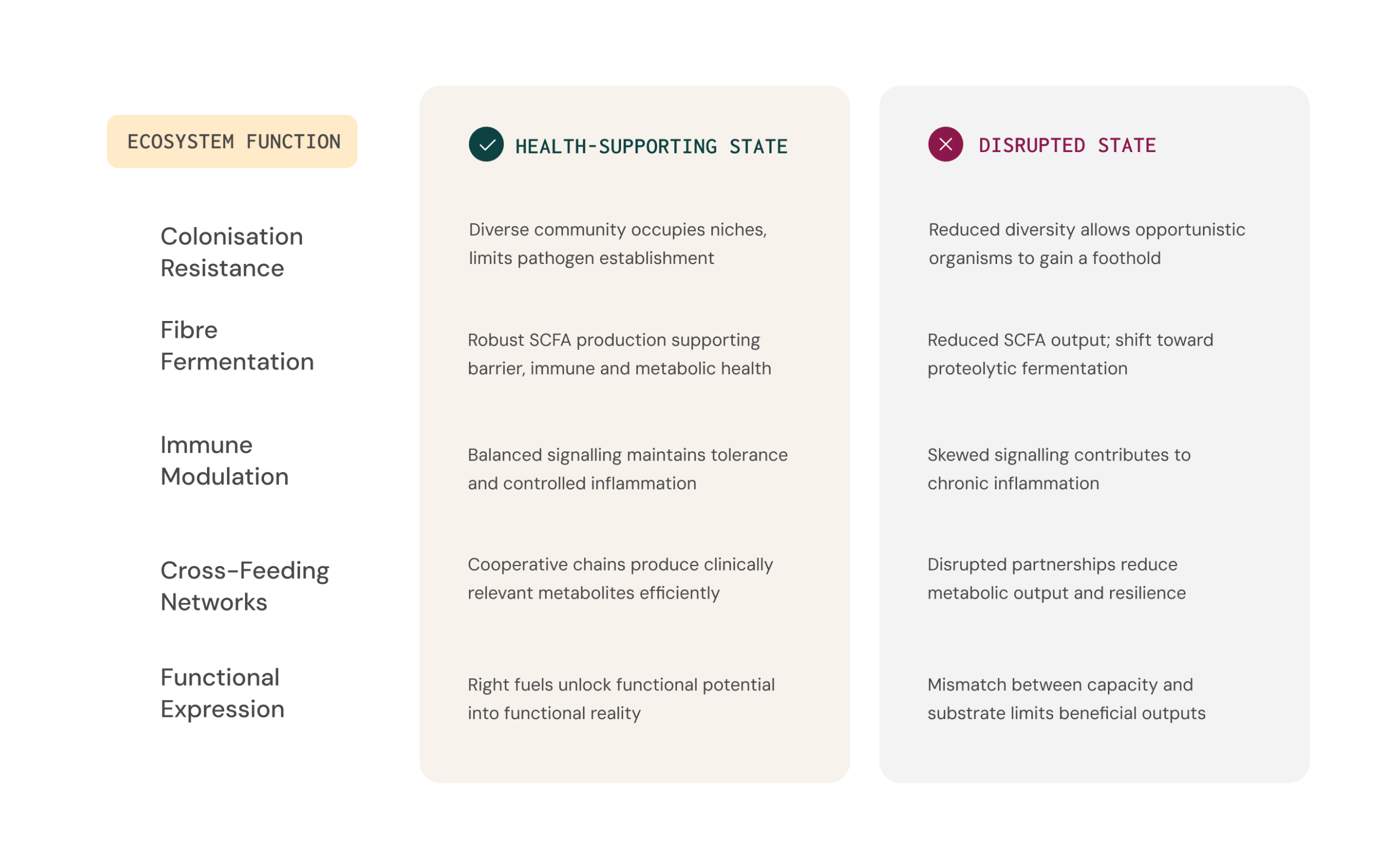
Health as a functional balance
Because healthy people can have very different species profiles, it is difficult to define a single "ideal" microbiome based only on who is present. It is more practical to think in terms of functional balance.
Functional balance
Functional dysbiosis
FUNCTIONAL BALANCE
SCFA production, immune regulation, barrier maintenance and colonisation resistance are robust
Pathways generating potentially harmful metabolites are kept in check.
FUNCTIONAL DYSBIOSIS
Beneficial activities are reduced and potentially harmful or symptom-linked functions are increased
Associated with distinct changes in metabolic functions linked to inflammation
A functional microbiome lens helps explain why the same intervention — higher fibre intake, probiotics or medication — may help one person but worsen symptoms in another, depending on microbial functions, dietary context and host factors. Supporting health often means working with the ecosystem: providing the right fuels, avoiding unnecessary disruptions, and using targeted strategies to shift microbial functions rather than chasing or eradicating individual species.

Key messages
Key POINT
The microbiome is a dynamic ecosystem whose functions — not just its composition — shape human health. Diverse communities support colonisation resistance, diet determines metabolic outputs, and health outcomes depend on community cooperation and functional balance rather than individual species.1,3
- Coyte KZ, Schluter J, Foster KR. The ecology of the microbiome: networks, competition, and stability. Science. 2015;350:663–666.
- Schroeder BO, Bäckhed F. Signals from the gut microbiota to distant organs in physiology and disease. Nat Med. 2016;22:1079–1089.
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–214.
- Sorbara MT, Pamer EG. Interbacterial mechanisms of colonization resistance and the strategies pathogens use to overcome them. Mucosal Immunol. 2019;12:1–9.
- Caballero-Flores G, Pickard JM, Núñez G. Microbiota-mediated colonization resistance: mechanisms and regulation. Nat Rev Microbiol. 2023;21:347–360.
- Kamada N, Chen GY, Inohara N, Núñez G. Control of pathogens and pathobionts by the gut microbiota. Nat Immunol. 2013;14:685–690.
- Buffie CG, Pamer EG. Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol. 2013;13:790–801.
- Jalanka J, Gunn D, Singh G, Krishnasamy S, Lingaya M, Crispie F, et al. Postinfective bowel dysfunction following Campylobacter enteritis is characterised by reduced microbiota diversity. Gut. 2023;72:451–459.
- Marshall JK, Thabane M, Garg AX, Clark WF, Moayyedi P, Collins SM. Eight year prognosis of postinfectious irritable bowel syndrome following waterborne bacterial dysentery. Gut. 2010;59:605–611.
- Rowland I, Gibson G, Heinken A, Scott K, Swann J, Thiele I, et al. Gut microbiota functions: metabolism of nutrients and other food components. Eur J Nutr. 2018;57:1–24.
- Koh A, De Vadder F, Kovatcheva-Datchary P, Bäckhed F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell. 2016;165:1332–1345.
- Teichmann J, Cockburn DW. In vitro fermentation reveals changes in butyrate production dependent on resistant starch source. Front Microbiol. 2021;12:640253.
- Venkataraman A, Sieber JR, Schmidt AW, Waldron C, Theis KR, Schmidt TM. Variable responses of human microbiomes to dietary supplementation with resistant starch. Microbiome. 2016;4:33.
- Ďásková N, Modos I, Krbcová M, Kuzma M, Pelantová H, Hradecký J, et al. Multi-omics signatures in new-onset diabetes predict metabolic response to dietary inulin. Nutr Diabetes. 2023;13:7.
- Alexeev EE, Lanis JM, Kao DJ, Campbell EL, Kelly CJ, Battista KD, et al. Microbiota-derived indole metabolites promote intestinal homeostasis through regulation of interleukin-10 receptor. Am J Pathol. 2018;188:1183–1194.
- Sinha AK, Laursen MF, Brinck JE, Rybtke ML, Hjørne AP, Procházková N, et al. Dietary fibre directs microbial tryptophan metabolism via metabolic interactions in the gut microbiota. Nat Microbiol. 2024;9:1964–1978.
- Peng L, Li ZR, Green RS, Holzman IR, Lin J. Butyrate enhances the intestinal barrier by facilitating tight junction assembly. J Nutr. 2009;139:1619–1625.
- Gilbert MS, Ijssennagger N, Kies AK, van Mil SWC. Protein fermentation in the gut; implications for intestinal dysfunction. Am J Physiol Gastrointest Liver Physiol. 2018;315:G159–G170.
- Corrêa-Oliveira R, Fachi JL, Vieira A, Sato FT, Vinolo MAR. Regulation of immune cell function by short-chain fatty acids. Clin Transl Immunol. 2016;5:e73.
- Canfora EE, Jocken JW, Blaak EE. Short-chain fatty acids in control of body weight and insulin sensitivity. Nat Rev Endocrinol. 2015;11:577–591.
- De Palma G, Shimbori C, Reed DE, Yu Y, Rabbia V, Lu J, et al. Histamine production by the gut microbiota induces visceral hyperalgesia through histamine 4 receptor signaling. Sci Transl Med. 2022;14:eabj1895.
- Gensollen T, Iyer SS, Kasper DL, Blumberg RS. How colonization by microbiota in early life shapes the immune system. Science. 2016;352:539–544.
- Wernroth M-L, Fall K, Svennblad B, Ludvigsson JF, Sjölander A, Almqvist C, et al. Early childhood antibiotic treatment for otitis media is associated with risk of type 1 diabetes. Diabetes Care. 2020;43:991–999.
- Kronman MP, Zaoutis TE, Haynes K, Feng R, Coffin SE. Antibiotic exposure and IBD development among children: a population-based cohort study. Pediatrics. 2012;130:e794–e803.
- Belkaid Y, Hand TW. Role of the microbiota in immunity and inflammation. Cell. 2014;157:121–141.
- Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, deRoos P, et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature. 2013;504:451–455.
- Zelante T, Iannitti RG, Cunha C, De Luca A, Giovannini G, Pieraccini G, et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity. Immunity. 2013;39:372–385.
- Mohr AE, Crawford M, Jasbi P, Fessler S, Sweazea KL. Lipopolysaccharide and the gut microbiota: considering structural variation. FEBS Lett. 2022;596:849–875.
- Zheng D, Liwinski T, Elinav E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020;30:492–506.
- Baruch EN, Youngster I, Ben-Betzalel G, Ortenberg R, Lahat A, Katz L, et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science. 2021;371:602–609.
- Culp EJ, Goodman AL. Cross-feeding in the gut microbiome: ecology and mechanisms. Cell Host Microbe. 2023;31:485–499.
- Palleja A, Mikkelsen KH, Forslund SK, Kashani A, Allin KH, Nielsen T, et al. Recovery of gut microbiota of healthy adults following antibiotic exposure. Nat Microbiol. 2018;3:1255–1265.
- Tiffany CR, Bäumler AJ. Dysbiosis: from fiction to function. Am J Physiol Gastrointest Liver Physiol. 2019;317:G602–G608.
- Chassard C, Dapoigny M, Scott KP, Crouzet L, Del’homme C, Marquet P, et al. Functional dysbiosis within the gut microbiota of patients with constipated-IBS. Aliment Pharmacol Ther. 2012;35:828–838.
- Tang WH, Wang Z, Levison BS, Koeth RA, Britt EB, Fu X, et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N Engl J Med. 2013;368:1575–1584.
- Lloyd-Price J, Arze C, Ananthakrishnan AN, Schirmer M, Avila-Pacheco J, Poon TW, et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature. 2019;569:655–662.