The gut microbiome
The human gut harbours trillions of microorganisms whose collective genome — the microbiome — encodes metabolic capabilities far exceeding those of the human genome alone.1 Over the past two decades, large-scale sequencing studies have demonstrated that the composition and function of this microbial community are associated with a wide range of health outcomes.2,3 Disruption of a healthy gut microbiome, often called dysbiosis, has been linked to gastrointestinal disorders, autoimmune conditions, cardiometabolic diseases, and neurological conditions.4
For healthcare practitioners, the clinical question is straightforward: can we measure a patient’s gut microbiome accurately enough, and interpret the results rigorously enough, to inform care?

Accurate gut measurement depends on getting the foundations right
MEASUREMENT TECHNOLOGY
The sequencing method used to capture the microbial community and its resolution, coverage, and accuracy
REFERENCE DATA QUALITY
The reference cohort against which results are compared and whether it eliminates technical and biological bias
SCIENTIFIC FRAMEWORK
The evidence curation applied to translate microbial signals into meaningful, clinically relevant context
Many gut health tests available today rely on older methods — culture-based assays, quantitative PCR (qPCR), or 16S ribosomal RNA (rRNA) gene sequencing — that provide only a partial view of the microbiome. These approaches can be good in specific circumstances (e.g. identification of pathogens), but have well-documented limitations in taxonomic resolution, coverage of the entire microbiome, and the ability to assess microbial function.6,7 The emergence of shotgun metagenomics — sequencing all DNA in a sample rather than a single gene — has fundamentally changed what is possible, enabling species- and even strain-level identification alongside functional pathway analysis.8
Microba has developed a comprehensive gut health test — Microba Microbiome Explorer — that combines accredited gastrointestinal diagnostics with high-resolution shotgun metagenomic microbiome profiling. The pathogen detection and gastrointestinal marker components are CE-certified and run within an ISO 15189 NATA-accredited medical laboratory.
Underpinning the test is a proprietary technology stack: a validated sample preservation system, a highly automated sequencing laboratory that has processed more than 100,000 metagenomes, and the Microba Community Profiler (MCP) — a peer-reviewed bioinformatic classifier that achieves highly accurate species identification in faecal samples.
100,000+
Metagenomes processed Operational track record
28,000+
Species assessed Species-level profiling
99%
Precision rate Virtually zero false positives
19,000+%
Consented profiles Internal validation dataset
Microba has developed a comprehensive gut health test — Microba Microbiome Explorer — that combines accredited gastrointestinal diagnostics with high-resolution shotgun metagenomic microbiome profiling.
The diagnostic components (pathogen detection and gastrointestinal markers) are CE-certified and run within an ISO 15189 NATA-accredited medical laboratory. The microbiome component is for research use only and provides species-level profiling and functional insights that go beyond what legacy testing methods can offer.
Underpinning the test is a proprietary technology stack: a validated sample preservation system, a highly automated sequencing laboratory that has processed more than 100,000 metagenomes, and the Microba Community Profiler (MCP) — a peer-reviewed bioinformatic classifier that achieves highly accurate species identification in faecal samples.
Did you know?
Microbial markers are scientifically curated using a rigorous three-tier evidence framework, and results are compared against a carefully defined healthy reference group of more than 450 individuals.
Validated sample preservation
protects the accuracy of every result
The accuracy of any microbiome test begins with sample integrity. Faecal samples are biologically active, and microbial composition can shift rapidly after collection if not properly preserved. Microba uses the Copan FLOQSwab in an active-drying tube (FLOQSwab-ADT) that has been rigorously benchmarked against commonly used preservation methods, including OMNIgene-GUT, RNAlater, BBL dry cotton swabs, and LifeGuard.9
In a peer-reviewed evaluation, Microba’s swab demonstrated the best performance for both technical (between-replicate) reproducibility and compositional stability relative to flash-frozen controls. Additionally, a second experiment in the same study demonstrated that the FLOQSwab-ADT maintained its performance across storage at -20°C, room temperature, and 50°C for four weeks, making it suitable for postal collection in a wide range of climatic conditions.9
Best reproducibility
Highest technical (between-replicate) reproducibility and compositional stability relative to flash-frozen controls in a peer-reviewed evaluation
Climate resilient
Stable across -20°C, room temperature, and 50°C for four weeks — suitable for postal collection Australia-wide
This means practitioners can be confident that the microbial profile generated from a swab collected at home closely reflects the true composition of the patient’s sample at the time of collection.
001
Patient collects
at home
002
FLOQSwab-ADT
preserves sample
003
Postal delivery in
any climate
004
Microba lab receives
intact sample
An ISO 15189 accredited laboratory has
processed over 100,000 metagenomes
Microba operates an ISO 15189 NATA-accredited medical testing laboratory — the internationally recognised standard for medical laboratories. The laboratory uses the latest DNA sequencing technology and maintains a high level of automation, from sample receipt through to data generation. An automated quality control pipeline monitors every sample, flagging any that fall outside predefined quality thresholds. To date, the laboratory has processed more than 100,000 metagenomes, providing a substantial operational track record.
How does it work?
The software systems used to analyse and interpret metagenomic data are developed under an ISO 13485 quality management system for software as a medical device. This dual accreditation — ISO 15189 for laboratory processes and ISO 13485 for software — reflects a commitment to clinical-grade quality at every stage.

ISO 15189
Internationally recognised standard for medical laboratory processes —
covering sample receipt, sequencing, and data generation
ISO 13485
Quality management system for software as a medical device —
covering the bioinformatic analysis and interpretation pipeline
Sequencing technology
Shotgun metagenomics identifies species that other methods miss entirely
Shotgun metagenomics sequences all DNA extracted from a faecal sample, providing a comprehensive and relatively unbiased view of the entire microbial community.8
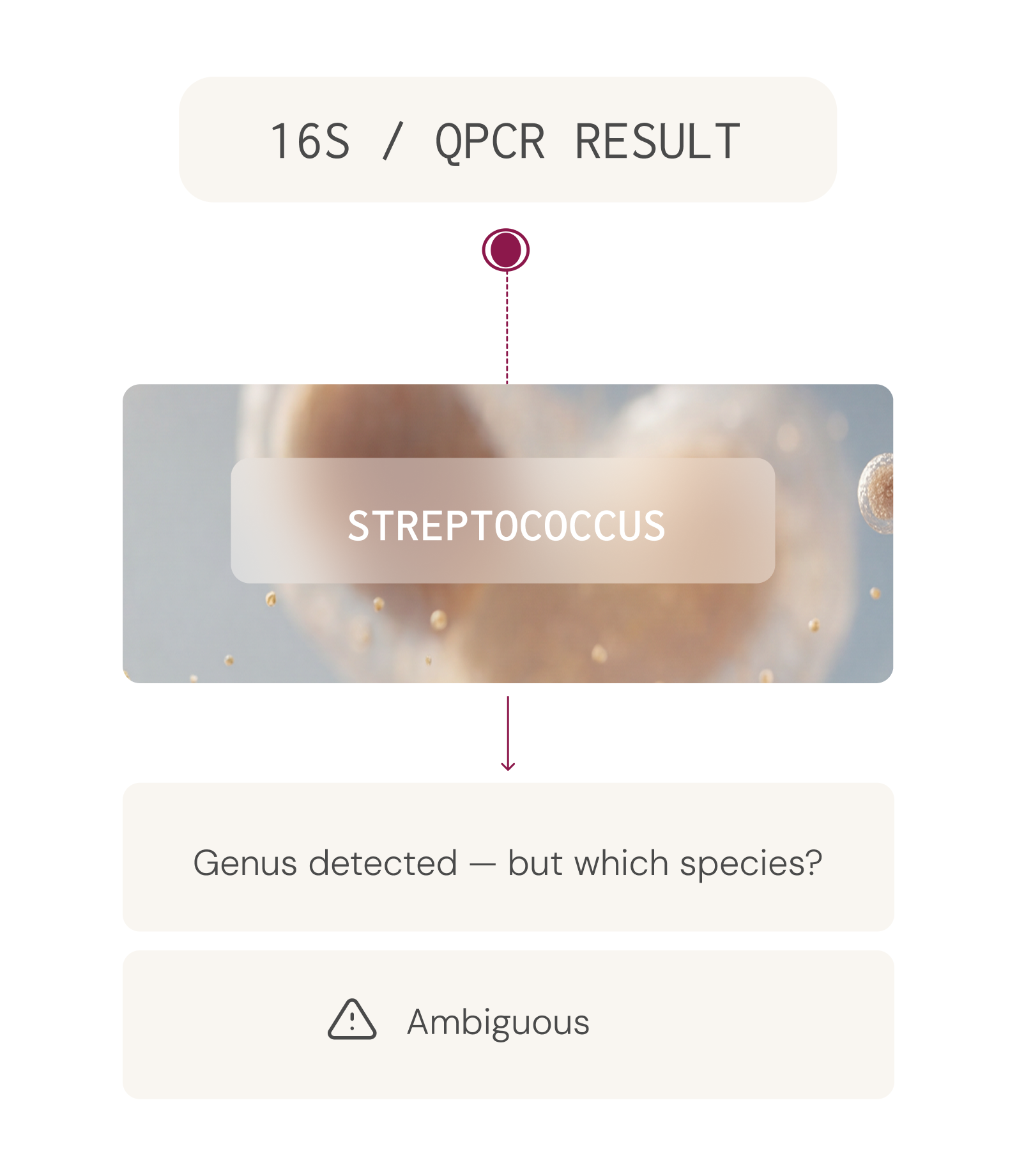
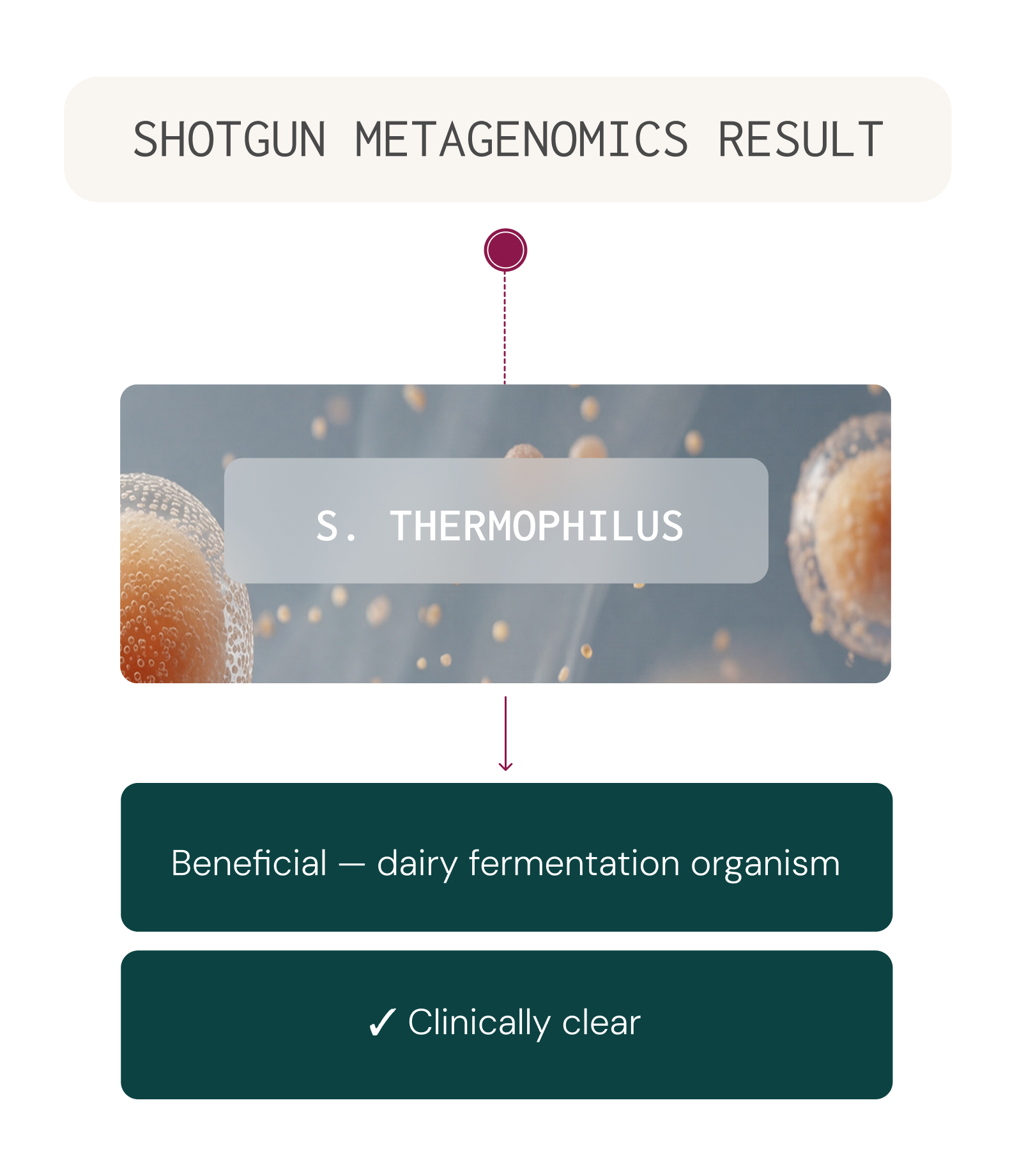
This allows identification of organisms at the species level — a critical capability, because different species within the same genus can have very different roles in health.
For example, Streptococcus thermophilus is a beneficial organism widely used in dairy fermentation, while Streptococcus anginosus is an opportunistic pathogen associated with abscess formation. A test that identifies only the genus Streptococcus cannot distinguish between these two very different clinical scenarios.
632 species vs 57
In a direct comparison, shotgun metagenomics identified 632 species in a sample where 16S sequencing detected only 57 — an order-of-magnitude difference in resolution that directly affects clinical utility.
Sequencing technology
Shotgun metagenomics
enables functional profiling
Equally important, shotgun metagenomics enables functional profiling — identifying the metabolic genes and pathways present across the entire microbial community. This makes it possible to assess the community’s capacity to produce health-relevant metabolites such as butyrate, or to carry out processes such as mucin degradation, providing a functional layer of information that 16S sequencing cannot accurately deliver.
18.96%
Butyrate-producing capacity detected across the microbial community — even when measured metabolite output appeared low.
Some tests reporting low short-chain fatty acid metabolites (measured via GC/MS) missed the broader picture. Microba’s community-level assessment showed 18.96% butyrate-producing capacity in the microbial community — indicating the functional machinery was present even when measured metabolite output appeared low. This shifts the clinical question from whether the microbiome is capable of producing butyrate to whether the right dietary substrates are available to support that production. 16S sequencing cannot deliver this functional layer of information — it identifies organisms but cannot assess which metabolic genes and pathways are present across the community.8

The Microba Community Profiler achieves 99% precision — peer-reviewed and benchmarked
Generating sequence data is only the first step. The accuracy of a microbiome profile depends critically
on the bioinformatic tools used to classify the millions of short DNA sequences (reads) produced by a sequencing run. The Microba Community Profiler (MCP) is a proprietary whole-genome alignment tool
that classifies metagenomic reads to produce species-level community profiles. Its performance has been rigorously benchmarked in a peer-reviewed study published in Frontiers in Microbiology, where it was evaluated against nine widely used academic classifiers across 140 simulated microbial communities of varying complexity.
PRECISION (% — HIGHER IS BETTER)
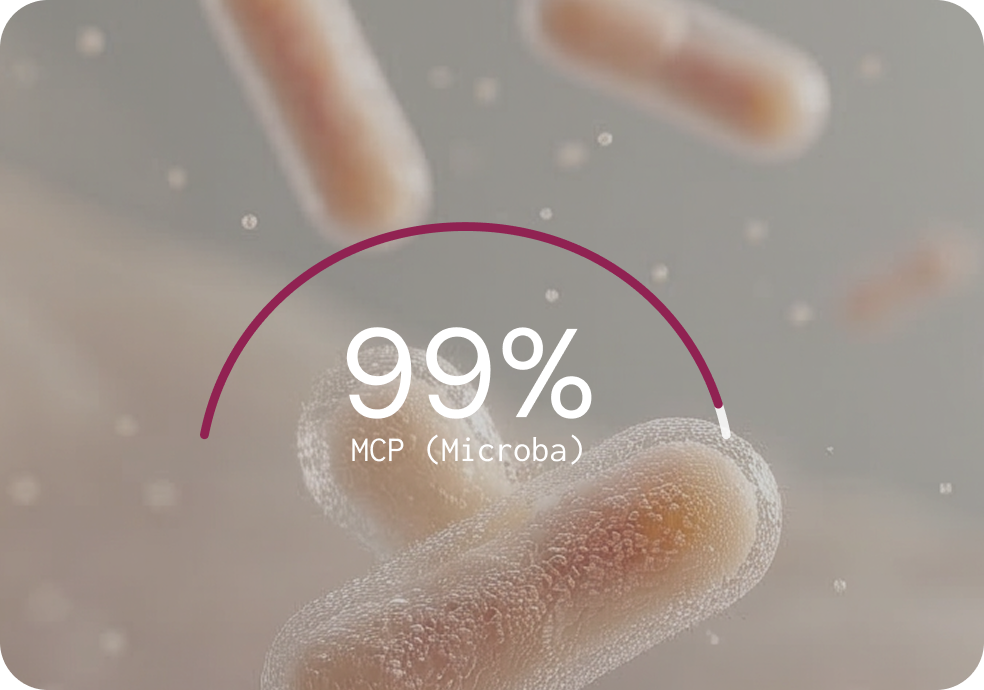
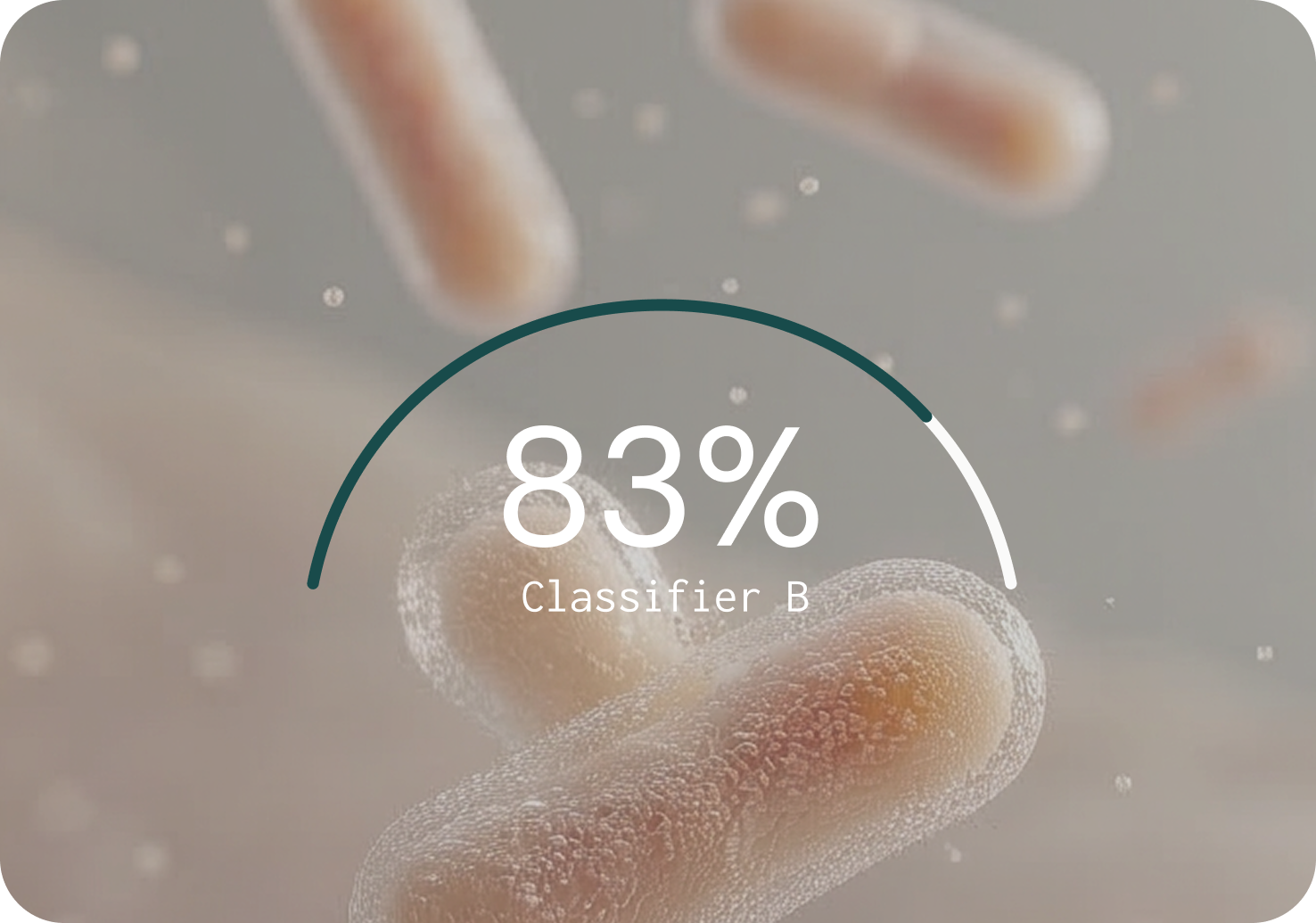
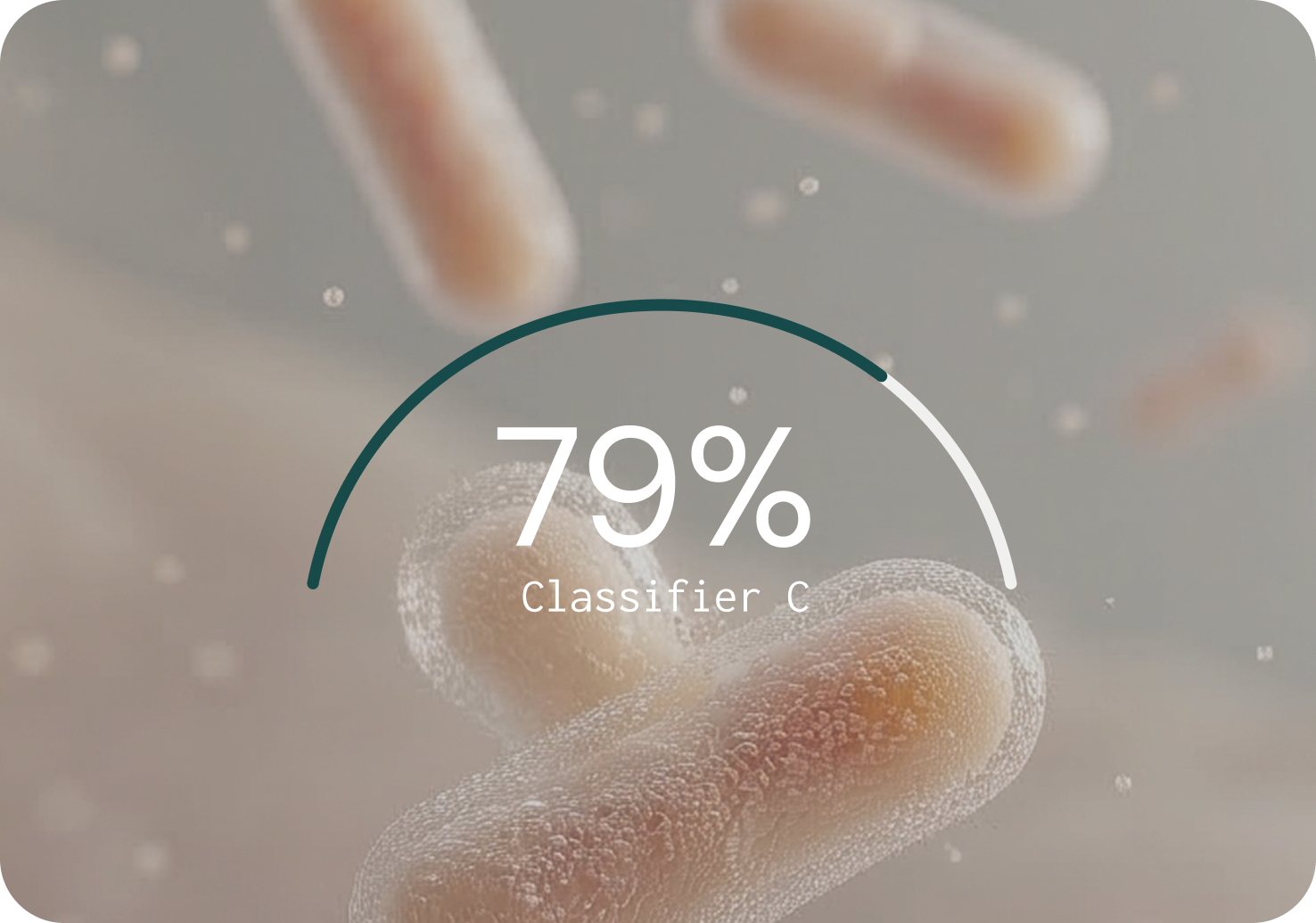
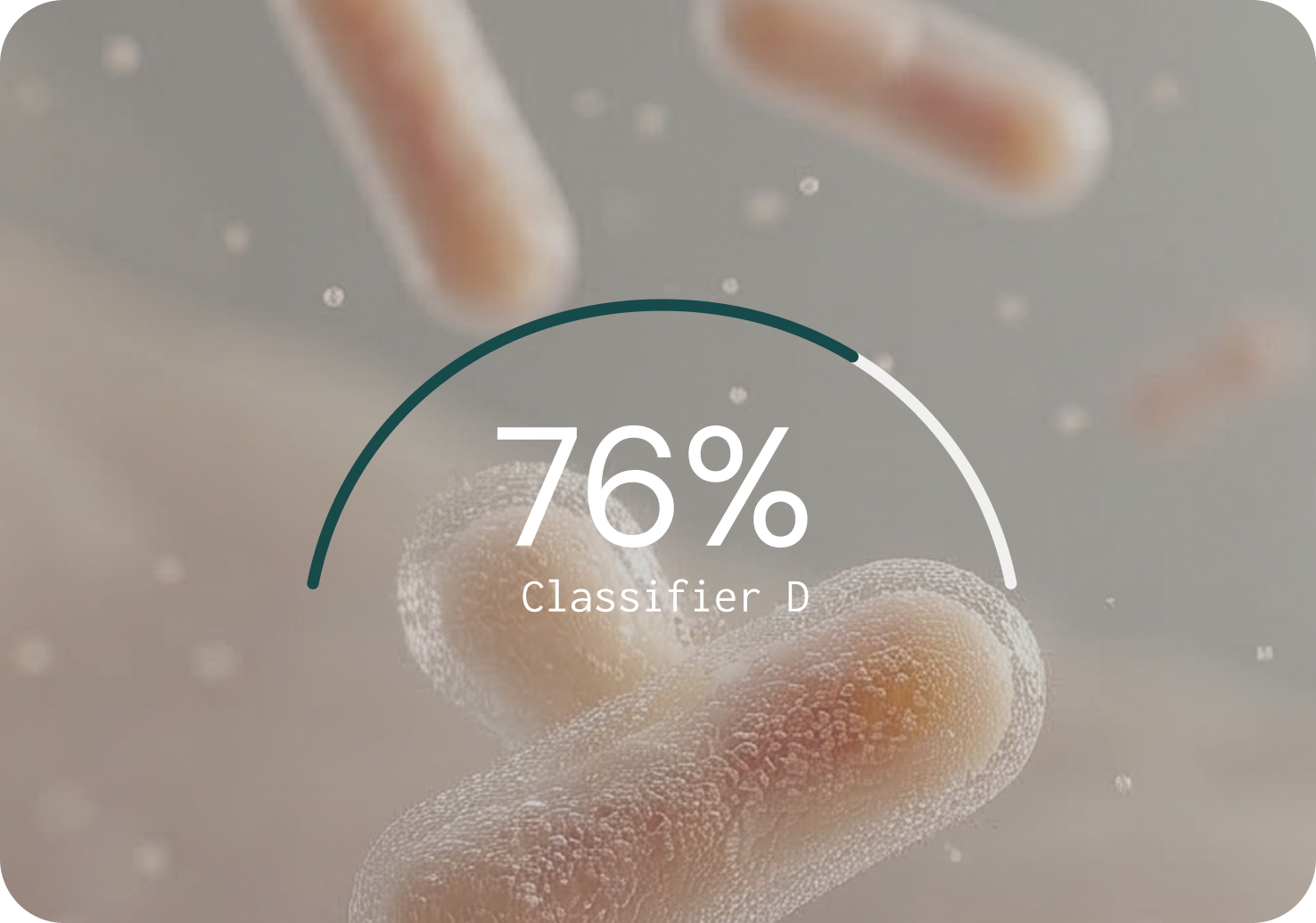
Key findings from the benchmarking study
A unique advantage of MCP is that it automatically filters species predictions to report only those with high confidence, without requiring the user to manually set abundance thresholds. Most other classifiers report thousands of low-abundance false positives unless the user manually inspects and filters the output.5
Three integrated layers connect microbial presence to measurable clinical markers
Microba Microbiome Explorer is a comprehensive gut health test that integrates three layers of information:
PATHOGEN DETECTION
A panel of 13 common bacterial pathogens and five parasites detected via CE-certified multiplex PCR assays.
HUMAN STOOL MARKERS
Six GI health markers including calprotectin, lactoferrin, faecal occult blood, secretory IgA, pancreatic elastase, and zonulin — assessed using CE-certified immunohistochemistry assays. Faecal pH is also measured as an investigative marker for research use only.
MICROBIOME PROFILING
Species-level profiling of 28,000+ species including microbial diversity, richness, and 16 health-associated functional markers — such as butyrate production, trimethylamine, hexa-acylated lipopolysaccharides, mucin degradation, and oxalate consumption. For research use only.
By combining diagnostic gastrointestinal markers with microbiome profiling, the test moves beyond simple species detection to provide a functional picture of gut health — linking what is present in the microbiome with measurable clinical markers of gut function and inflammation.
Only markers passing all three evidence tiers are included in the report
Not every microbial signal is clinically meaningful. To ensure that the microbiome markers reported in the test are supported by robust evidence, Microba applies a rigorous three-tier scientific curation framework. Only markers that satisfy all three tiers (with limited, clearly disclosed exceptions) are included in the report.*
*Exceptions in Microbiome Explorer: acetate and intestinal inflammation (mechanism only), beta-glucuronidase and impaired detoxification (mechanism only) and Emerging Markers.”
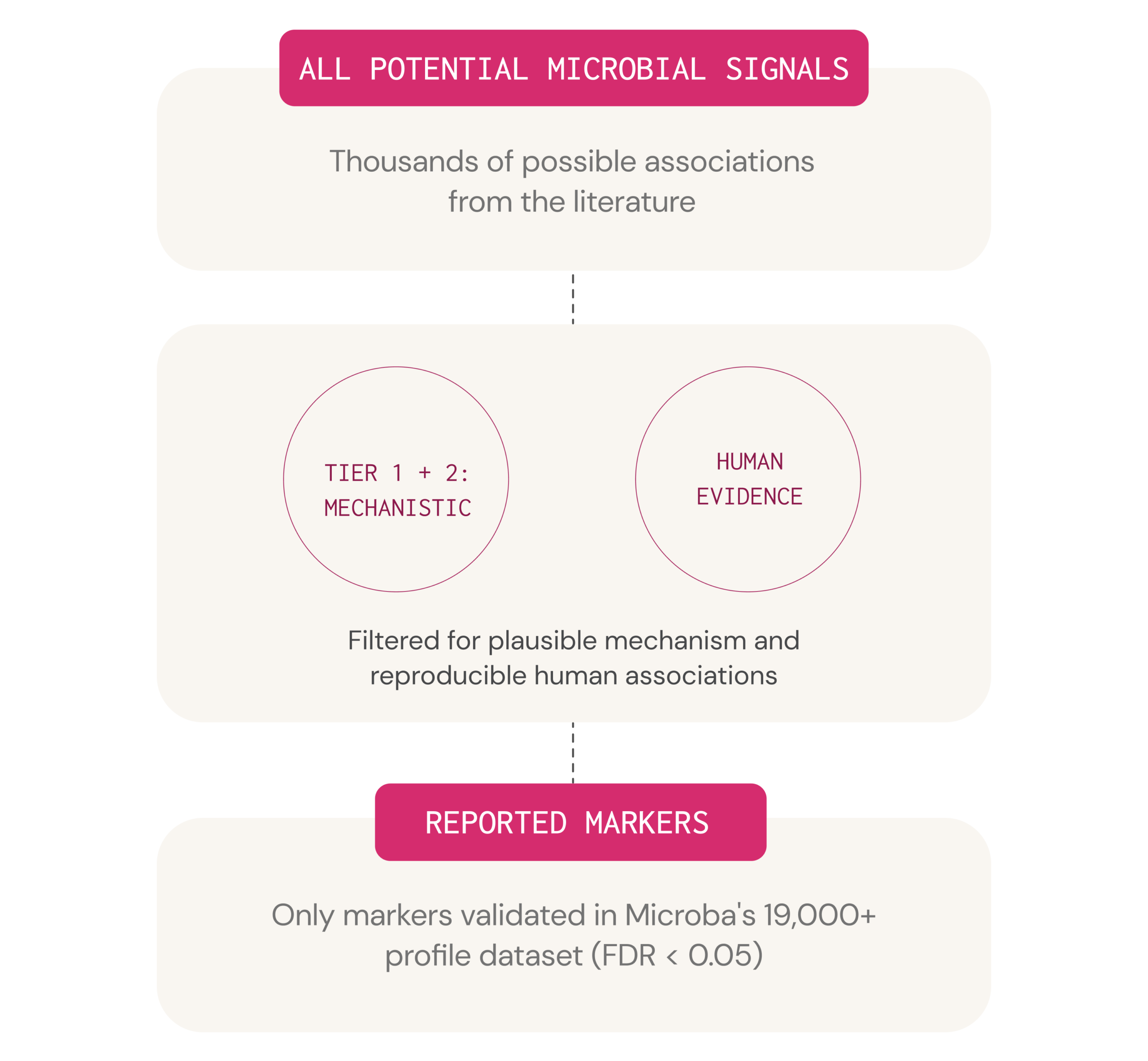

Plausible mechanism of action
SCFA production, immune regulation, barrier maintenance and colonisation resistance are robust. Pathways generating potentially harmful metabolites are kept in check.
Example: The link between mucin-degrading species and intestinal inflammation is supported by mechanistic studies showing that depletion of the mucus barrier increases microbial proximity to the epithelium, driving immune activation.10,13
Reproducible human associations
At least two peer-reviewed human studies must demonstrate a direct or indirect association between the microbial marker and the health outcome.
Example: Direct evidence includes correlation of the marker with a clinical measure, such as oral species abundance and faecal calprotectin. Indirect evidence includes correlation with relevant diseases, or plasma levels of the microbial marker with a clinical measure.14,15
Significant associations in Microba’s dataset
The microbial marker must demonstrate a statistically significant association with a minimum small effect size in Microba’s database of more than 19,000 consented metagenomic profiles with linked health and lifestyle data. Analyses are controlled for age, sex, BMI, and Bristol stool index.
Example: This internal validation step ensures that markers are not only supported by published literature but are also reproducible in a large, independent cohort processed through Microba’s own laboratory and analysis pipeline.
Reference data
A carefully defined healthy reference group of 450+ individuals eliminates technical bias
To determine whether a patient’s microbiome markers are within a healthy range, results must be compared against an appropriate reference group. Many commercially available tests either provide no details about their reference cohort, use publicly available microbiome data (which introduces significant variability due to differences in sample collection, processing, and analysis methods), or compare samples against their entire database regardless of health status.
Microba’s healthy reference group has been carefully selected and includes more than 450 individuals who meet strict inclusion criteria. Critically, all reference samples were collected and processed using exactly the same workflow as patient test samples, eliminating a common source of technical bias.
INCLUSION CRITERIA
No major medical conditions
No or minimal GI symptoms· Mild or lower stress, anxiety, and depression
BMI below 30 Daily fruit and vegetable intake
Low to moderate alcohol consumption
Evidence-graded actions give practitioners
a clear basis for clinical conversations
Generating accurate microbiome data is necessary but not sufficient. For gut health testing to be clinically useful, the data must be translated into interpretable insights that healthcare practitioners can act upon.
In Microba’s Microbiome Explorer, microbial markers and gastrointestinal markers are organised into health categories that correspond to recognisable clinical concepts. Each health category provides a clear interpretation of whether the patient’s results fall within or outside the healthy reference range, and, where applicable, links to the relevant diagnostic GI markers for clinical correlation.
When a marker is identified as out of range, the report provides evidence-graded possible actions. Microba’s science team has undertaken a rigorous review of the available scientific evidence for different dietary, supplement, or lifestyle interventions to modulate microbial functions and graded these using the NHMRC evidence grading framework. Each listed action is accompanied by the evidence grade and hyperlinks to the references that informed the grade.
Mucin degradation and intestinal
inflammation
The proportion of mucin-degrading species in the microbiome is one of the markers assessed in the test. The mechanistic basis for this marker is well established: when dietary fibre is insufficient, mucin-degrading microbes consume the protective mucus layer lining the gut, increasing microbial contact with the intestinal epithelium and triggering immune activation.
A large cross-sectional study of more than 1,000 individuals found a significant positive association between mucin-degrading pathway abundance and faecal calprotectin (a clinical marker of intestinal inflammation). Elevated mucin degrading pathways have also been observed in colorectal cancer cohorts.
In Microba’s own dataset, the relative abundance of mucin-degrading species is significantly increased in several health conditions related to intestinal inflammation compared to healthy controls, after controlling for confounders.
This layered approach — mechanistic plausibility, published human evidence, and internal validation — gives practitioners a clear rationale for each marker and a basis for evidence-informed clinical conversations with patients. Microba Microbiome Explorer is exclusively offered through healthcare professionals. This distribution model ensures that results are interpreted accurately, responsibly, and in the best interests of patient care.
What makes Microba’s
approach different
The gut health testing market includes a range of products built on different technologies, evidence standards, and quality systems. Several features distinguish Microba’s approach from legacy and competing tests.
not 16S or qPCR
benchmarked bioinformatics
evidence curation
reference group
Transparency about limitations is essential for responsible clinical use
The microbiome component of Microba Microbiome Explorer is for research use only and is not a diagnostic tool. Microbiome results should be interpreted by qualified healthcare practitioners in the context of a patient’s clinical history, symptoms, and other diagnostic findings.
The test does not diagnose, treat, or prevent any disease. Like all sequencing-based microbiome tests, Microba’s metagenomic profiling produces compositional data — it measures the relative proportions of different species rather than their absolute quantities. This is an inherent characteristic of current sequencing technology and means that changes in the abundance of one organism can influence the apparent abundance of others.
Microba's reference database, while comprehensive, does not capture the full diversity of the human gut microbiome.
On average, Microba will be able to assign a species ID to 82.4% of sequencing reads. Microba continues to expand the database with newly identified species and strains, and ongoing updates to the reference database aim to increase the proportion of sequencing reads assigned a species ID and lower detection limits further.
The marker curation framework is also a living process: as new evidence emerges, markers may be added, refined, or retired. Future directions include further validation studies, expansion of the healthy reference group to include additional populations, and continued development of evidence-graded suggested actions informed by the latest intervention research.
A considered integration of science,
evidence, and quality
Microba’s Microbiome Explorer represents a considered integration of advanced metagenomic science, rigorous evidence curation, and accredited laboratory quality. By combining CE-certified gastrointestinal diagnostics with high-resolution microbiome profiling, the test provides healthcare practitioners with a comprehensive view of gut health that goes well beyond what legacy testing methods can offer.
Every layer of the technology stack — from validated sample preservation and automated laboratory processing, through to the peer-reviewed MCP — has been designed to maximise accuracy and minimise the risk of misleading results. The three-tier marker curation framework ensures that only microbial markers with robust supporting evidence are included, and results are compared against a carefully curated healthy reference group.
Microba commitment
While the microbiome component remains for research use only, the depth and quality of information it provides can support practitioners in building a more complete understanding of their patients’ gastrointestinal health.
As the science of the microbiome continues to advance, Microba is committed to translating that science into testing that practitioners can trust.
Sample collection
- Validated FLOQSwab-ADT
- Climate-resilient postal collection
Lab + data generation
- ISO 15189 accredited lab
- 100,000+ metagenomes
- Automated QC pipeline
Core analysis (the “engine”)
- Shotgun metagenomics
- Microba Community Profiler (99% precision)
Interpretation layer
- Three-tier evidence curation
- 450+ healthy reference group
- NHMRC-graded actions
Final output
- Clinically useful insights for practitioners
. Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016). https://doi.org/10.1056/NEJMra1600266
2. Gilbert, J. A., Blaser, M. J., Caporaso, J. G., Jansson, J. K., Lynch, S. V. & Knight, R. Current understanding of the human microbiome. Nat. Med. 24, 392–400 (2018). https://doi.org/10.1038/nm.4517
3. Fan, Y. & Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 19, 55–71 (2021). https://doi.org/10.1038/s41579-020-0433-9
4. Zmora, N., Suez, J. & Elinav, E. You are what you eat: diet, health and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 16, 35–56 (2019). https://doi.org/10.1038/s41575-018-0061-2
5. Parks, D. H., Rigato, F., Vera-Wolf, P., Krause, L., Hugenholtz, P., Tyson, G. W. & Wood, D. L. A. Evaluation of the Microba Community Profiler for taxonomic profiling of metagenomic datasets from the human gut microbiome. Front. Microbiol. 12, 643682 (2021). https://doi.org/10.3389/fmicb.2021.643682
6. Jovel, J., Patterson, J., Wang, W., Hotte, N., O’Keefe, S., Mitchel, T. et al. Characterization of the gut microbiome using 16S or shotgun metagenomics. Front. Microbiol. 1591-019-0406-6
7, 459 (2016). https://doi.org/10.3389/fmicb.2016.00459
7. Sczyrba, A., Hofmann, P., Belmann, P., Koslicki, D., Janssen, S., Dröge, J. et al. Critical assessment of metagenome interpretation — a benchmark of metagenomics software. Nat. Methods 14, 1063–1071 (2017). https://doi.org/10.1038/nmeth.4458
8. Hugenholtz, P. & Tyson, G. W. Metagenomics. Nature 455, 481–483 (2008). https://doi.org/10.1038/455481a