Why it matters for interpreting microbiome balance
A healthy gut microbiome is a balanced one. Evaluating that balance cannot be reduced to a list of well-known species. It reflects the overall state of the ecosystem shaped by diversity, relative abundance, and the functional capacity of the microbial community.
Many testing approaches still measure only fragments of this system – selected pathogens, limited taxa, or small gene regions and attempt to infer balance from that partial view.
If the clinical question is ecosystem-level balance, the measurement must reflect the complexity of the ecosystem.
Species-level detection
Measures the presence or absence of individual organisms in isolation - no diversity data, no functional insight, no understanding of how the community behaves as a whole.
Ecosystem-level assessment
Captures the full community picture: diversity, relative abundance, functional capacity, and the physiological conditions that shape how the microbiome actually functions.
Partial testing can’t reliably answer critical questions
In clinical practice, microbiome testing is rarely about detection alone.
The goal is interpretation: understanding how microbial ecology may be influencing:

Chronic symptoms

Inflammation
Gut barrier integrity

Metabolic signalling
That interpretation requires context.
A partial view of the microbiome may detect something, but it can’t reliably answer the more clinically meaningful questions
How does this organism’s abundance compare to the overall ecosystem? How does this organism’s abundance compare to the overall ecosystem?
Is the ecosystem resilient or vulnerable to instability?
Is functional capacity reduced?
Is it contributing to an anti-inflammatory or pro-inflammatory state?
Microbiome balance emerges from the whole system, targeted tests are inherently limited in what they can explain. Healthy gut microbial communities are characterised by high taxa diversity, high microbial gene richness, and a stable functional core, properties that cannot be captured by targeted detection methods alone.¹
Testing technologies vary in what they can measure and reveal
Microbiome testing has progressed through several technological stages. Each has improved capability, but not all are designed to assess ecosystem-level balance. Understanding these differences helps clinicians choose appropriate tools and interpret results correctly.
Culture-Based Testing Useful for pathogen detection, not for ecosystem assessment. Culture remains valuable for identifying certain infectious organisms and guiding antimicrobial therapy2. However: Many gut organisms cannot be cultured using standard laboratory techniques. Culture favours organisms that grow well in-vitro, not necessarily those most relevant in-vivo. It provides no information about diversity or ecological relationships.
Highly sensitive - but only for selected targets. PCR-based testing is extremely useful when there is a specific clinical hypothesis (e.g., presence of potential pathogens such as C. difficile, Giardia).2 However:2 You can only detect what you test for. It does not provide ecosystem-level composition. It does not assess diversity or resilience. It does not evaluate functional capacity. It cannot identify unexpected or emerging organisms. PCR answers specific questions well, however it is not designed to assess ecosystem balance.
Broader view, but limited resolution and no direct functional insight. 16S rRNA gene sequencing (16S) allows a broader survey of bacterial communities without culture. However, 16S rRNA sequencing is an amplicon-based method. It targets and amplifies a small region of the 16S ribosomal RNA gene, which is only present in bacteria and Archaea. This creates several clinically relevant limitations: Often limited to genus-level resolution, with species-level identification carrying a high rate of false positives³ ⁴ Inability to reliably distinguish clinically important species within the same genus. No detection of fungi or protists and 16S rRNA targets often miss Archaea³ ⁴ It cannot identify novel or uncharacterised organisms No direct measurement of microbial functional genes or pathways. Research directly comparing the two approaches confirms that 16S detects only part of the gut microbiota community revealed by shotgun sequencing, and that less abundant — but biologically meaningful, taxa are largely invisible to 16S methods.⁴ 16S can reveal broad trends, but it does not provide the resolution or functional depth required for high-confidence clinical interpretation.
Whole-community, species-level, function-aware measurement. Shotgun metagenomics sequences all DNA in a sample. This enables clinicians to assess: Composition: species and even strain-level identification of microorganisms Breadth: bacteria, fungi, Aarchaea, protists and other microbial DNA Discovery: detection of previously uncharacterised organisms Function: microbial genes and metabolic pathways Studies demonstrate that shotgun metagenomicwhole genome sequencing has multiple advantages over 16S amplicon methods, including enhanced detection of bacterial species, increased detection of diversity, and direct assessment of increased predictionfunctional of genes.5 6
The critical shift
Metagenomics enables ecosystem-level assessment. Because it captures the full genetic content of the community. ⁵ Not just selected fragments
Comparison of microbiome testing methods
The table below summarises how common microbiome testing approaches differ in scope, resolution, and suitability for assessing ecosystem-level balance.
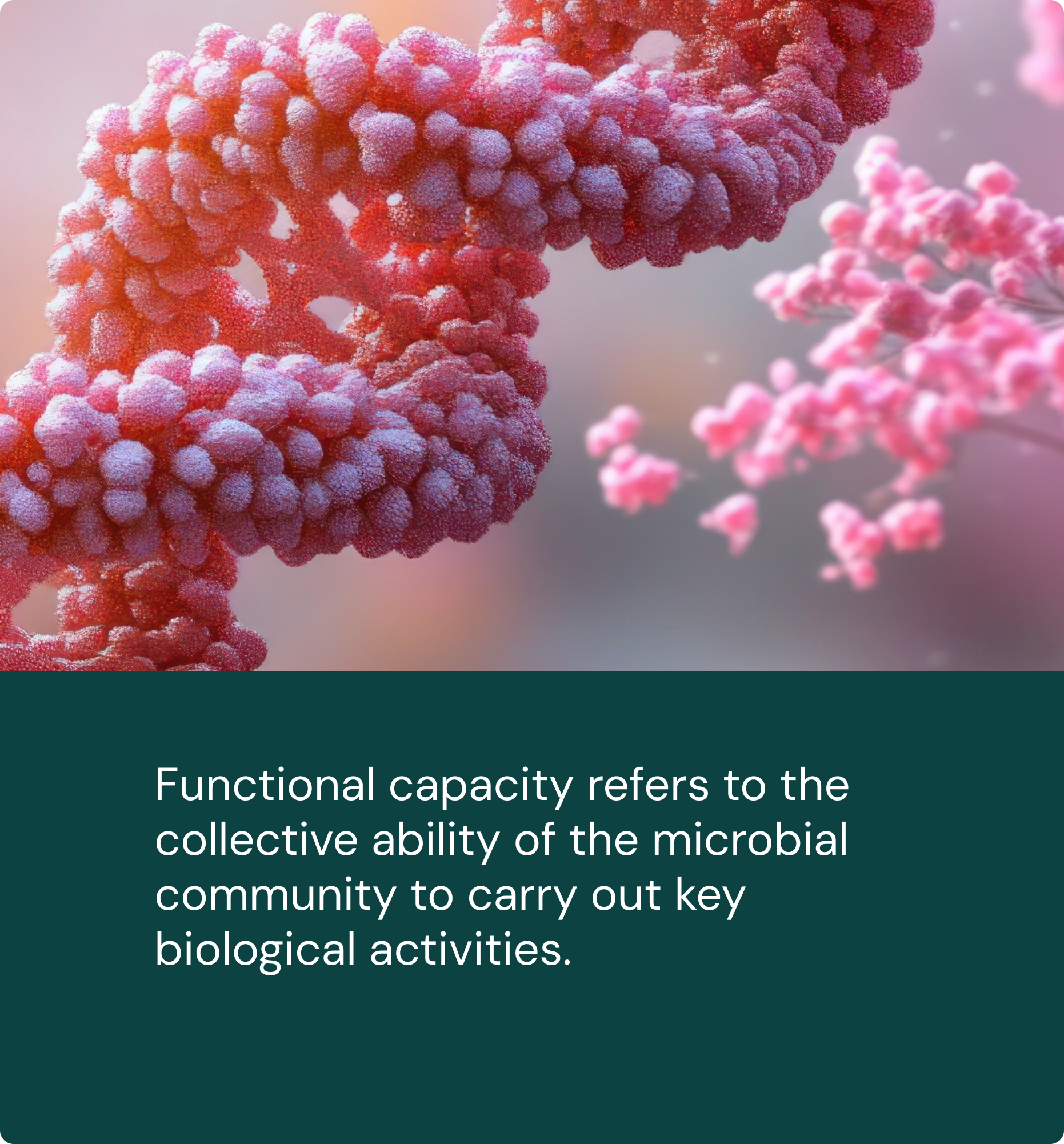
Why functional capacity matters
In an ecological model, clinical impact is determined not just by which organisms are present, but by what the ecosystem can do.
This is called the Functional capacity. These functions are distributed across multiple organisms that share metabolic pathways.
Clinically relevant examples include
Short-chain fatty acid (SCFA) production (e.g. butyrate)
SCFAs are produced through bacterial fermentation of dietary fibre and play key roles in gut barrier integrity, immune modulation, and inflammation control. Reduced SCFA-producing capacity has been linked to increased intestinal permeability and systemic inflammatory burden.⁷ ⁸
Pro-inflammatory LPS signalling potential
LPS produced by Gram-negative bacteria triggers innate immune activation. Dysbiosis, with reduced diversity and overgrowth of pro-inflammatory LPS-producing organisms, shifts the microbiome towards a pro-inflammatory state, contributing to low-grade systemic inflammation and chronic disease risk.⁹ ¹⁰
Mucin degradation activity
Mucin-degrading organisms including Akkermansia muciniphila and Bacteroides species can promote mucus production when at optimal levels but can deplete the mucus layer at high levels. Excessive mucus degradation can compromise mucosal integrity and increase immune activation. ¹¹ ¹²
Protein fermentation by-products
Fermentation of undigested protein in the distal colon produces metabolites including trimethylamine, hydrogen sulphide, branched-chain amino acids, ammonia and phenolic compounds, some of which drive intestinal inflammation and may contribute to systemic toxic load at elevated concentrations.
Without functional information
Clinicians are left interpreting composition alone which often fails to explain chronic symptoms, relapse cycles, or systemic inflammatory burden.
Whole-ecosystem testing changes the clinical questions
When clinicians move from targeted detection to whole-community measurement, the clinical questions evolve.
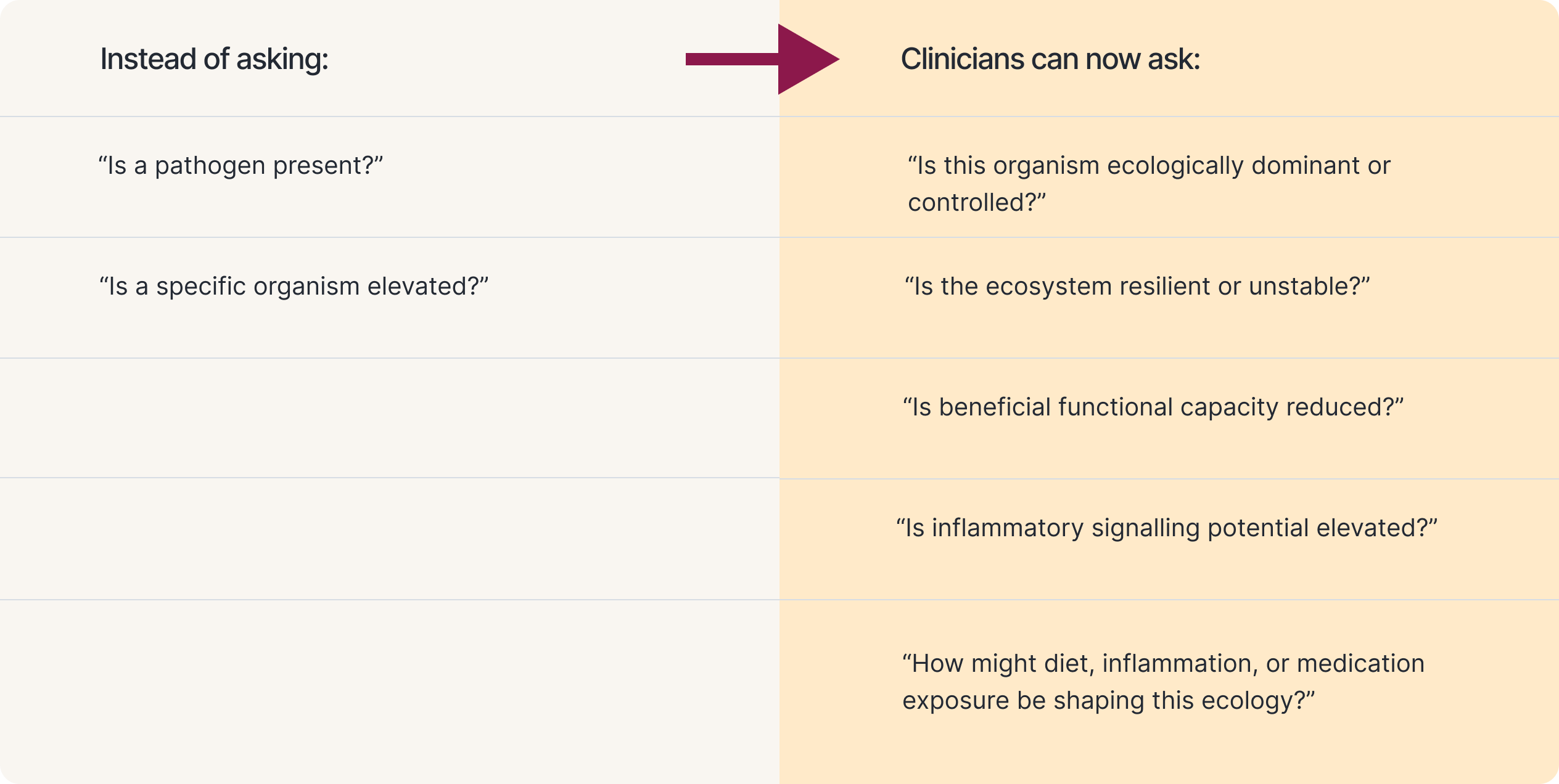

These are the questions that matter in complex, chronic presentations. Research linking gut dysbiosis to over 117 gastrointestinal and extra-gastrointestinal diseases underscore why understanding the ecosystem state, not just species presence, is clinically essential.¹³
What makes microbiome testing clinical-grade?
Whole-community sequencing is a necessary foundation, but clinical application requires more.
For metagenomics to be clinically useful, it must be delivered through validated and accredited processes. Clinical-grade testing requires:

Validated sample collection and preservation
The microbiome changes rapidly once a sample is produced. Without robust preservation, results can be distorted by microbial growth, degradation, or DNA fragmentation.

Accredited laboratory standards (e.g., ISO 15189)
Clinical-grade testing requires operation to medical laboratory standards, including quality systems, traceability, and analytical validation.

High-quality bioinformatics pipelines
Metagenomic data is only as useful as the computational system interpreting it. Low-quality bioinformatics increases false positives, misclassification, and missing data.
Evidence-based interpretation frameworks
Results must be aligned with reproducible evidence in humans and presented in a clinically usable format.
Metagenomics is only clinically meaningful when results are accurate, reproducible, and interpretable.
A Practical Summary for Clinicians
Microbiome balance is an ecosystem property. It cannot be accurately assessed through partial or targeted testing alone.
Culture, PCR, and 16S methods can answer specific questions but they do not provide whole-ecosystem visibility.
If the clinical goal is ecosystem-level understanding, whole-community metagenomics is required.
And if the goal is actionable interpretation, the metagenomic assessments must be delivered through validated, clinical-grade systems.