What is the
gut microbiome
The gut microbiome is the community of microorganisms, together with their genes and metabolic functions, that live in the gastrointestinal tract. While microbes exist across multiple body sites, the gut microbiome is the most extensively studied because of its density, diversity and metabolic activity. The gut microbiome is best understood as an ecosystem: a living community of many different microbes that interact with each other and with the human body.
The gut microbiome is
a measurable biological system
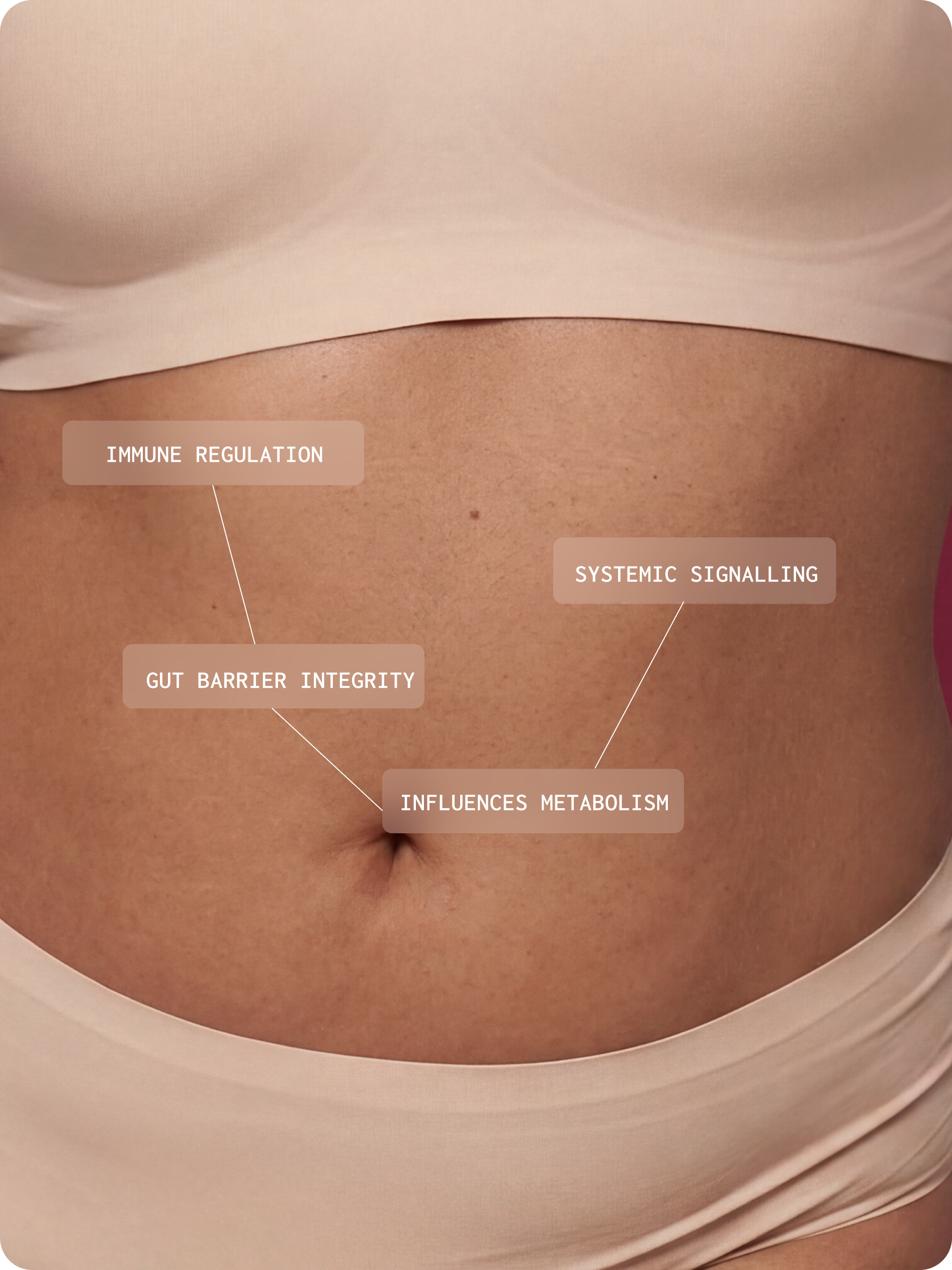
The gut microbiome is a dynamic biological system that is essential to normal human physiology, influencing immune regulation, gut barrier integrity, metabolism, and systemic signaling.1
It is a clinically relevant biological system with meaningful implications for complex and multi-system presentations.
The healthy function of the microbiome depends on a balanced ecosystem
The nature of these interactions is not fixed, it depends on the gut environment and overall ecological balance. These interactions generally fall into four categories
Bacteria
Archaea
Viruses
Fungi
Microbial interactions are shaped by balance, not fixed labels
The nature of these interactions is not fixed, it depends on the gut environment and overall ecological balance. These interactions generally fall into four categories:
Present without measurable benefit or harm to the host.
Contributes positively to metabolism, immune regulation or barrier integrity.
Typically neutral, but may cause harm when ecological balance shifts.
Capable of causing disease or disrupting normal function.
Microbes convert fuel into compounds that influence health
Gut microbes feed on available substrates including dietary fibre, proteins and host-derived compounds and convert them into metabolites that interact with body systems.1,2 What they produce depends on which microbes are present, the functional pathways they carry, and the fuel sources available to them.
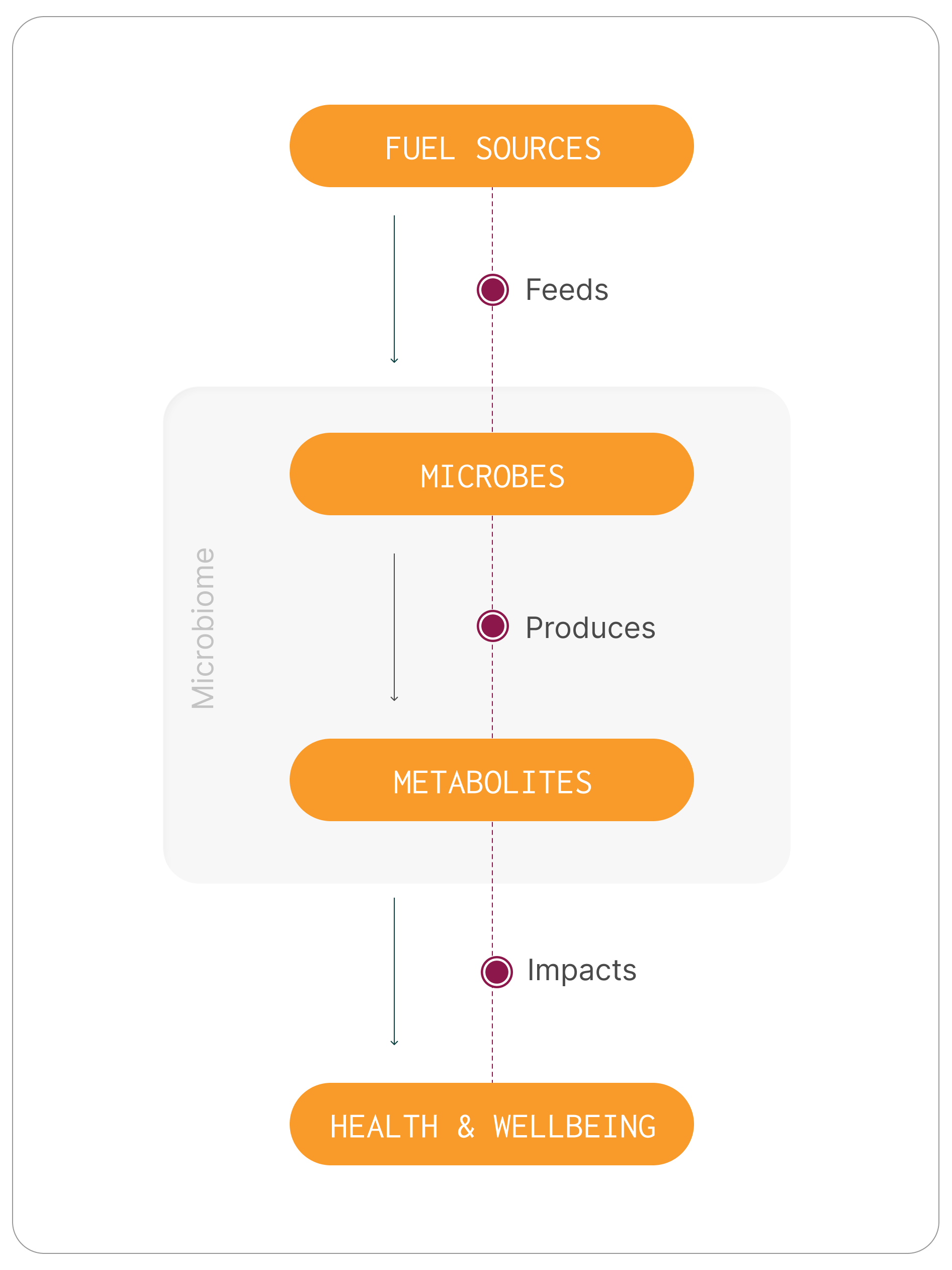
Key helpful functions of the gut microbiome
Microbes influence human physiology through the compounds they produce and the signals they generate. These microbial metabolites interact with immune pathways, support gut barrier integrity, and contribute to broader metabolic regulation.
Metabolism
Microbial metabolites influence energy use, lipid processing and systemic metabolic balance
Immune regulation
The gut microbiome helps regulate immune responses and maintain immune balance
Gut barrier function
Microbial activity supports epithelial integrity
Gut–brain & systemic signalling
Microbial activity supports epithelial integrity
The microbiome functions as part of human biology — not separate from it.
It’s not enough to know which microbes are present, you need to know what the ecosystem is doing
Microbiome species vary widely between individuals, yet different species can produce the same essential metabolites. Two people with very different microbial profiles may share comparable functional output.4-6
This is why clinical insight increasingly focuses on metabolic activity, such as butyrate production, rather than species detection alone..

Microbiome insight adds a systems-level lens to clinical care.
The microbiome interacts with multiple biological systems simultaneously, it can provide additional interpretive insight in chronic and complex cases.
In multi-system presentations not fully explained by traditional markers, understanding microbial function adds a broader systems-level perspective.
In multi-system presentations not fully explained by traditional markers, understanding microbial function adds a broader systems-level perspective.
In multi-system presentations not fully explained by traditional markers, understanding microbial function adds a broader systems-level perspective.
Ecological balance
Functional pathways
Metabolic output
Patient context
Key Takeaway
The human microbiome is a measurable biological system that functions as part of normal human physiology.
Community-wide measures such as diversity and functional output can be more clinically meaningful than detection of specific species.
As understanding of the microbiome grows, it is becoming an increasingly valuable part of clinical assessment, especially in chronic or multi-system presentations.
References:
- Sender R. Fuchs S. & Milo R. Revised Estimates for the Number of Human and Bacteria Cells in the Body.PLoSBiology 14 e1002533 (2016).
- HuttenhowerC. et al. Structure function and diversity of the healthy human microbiome. Nature 486 207–214 (2012).
- Jiang Y. Che L. & Li S. C. Deciphering the personalized functional redundancy hierarchy in the gut microbiome. Microbiome 14 17 (2025).
- Fan Y. & Pedersen O. Gut microbiota in human metabolic health and disease. Nat. Rev.Microbiol. 19 55–71 (2021).
- Lynch S. V. & Pedersen O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375 2369–2379 (2016).
- Koh A. DeVadderF. Kovatcheva-Datchary P. & Bäckhed F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 165 1332–1345 (2016).
- ZhernakovaA. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352 565–569 (2016).
- De Filippis F. et al. Distinct genetic and functional traits of human intestinalPrevotellacopri strains are associated with different habitual diets. Cell Host Microbe 25 444–453 (2019).
- Zhang Z. J. et al.Comprehensive analyses of a large human gutBacteroidales culture collection reveal species- and strain-level diversity and evolution. Cell Host Microbe 32 1853–1867 (2024).
- Tian L. et al. Deciphering functional redundancy in the human microbiome. Nat Commun 11 6217 (2020).
- denBestenG. et al. The role of short-chain fatty acids in the interplay between diet gut microbiota and host energy metabolism. Journal of Lipid Research 54 2325–2340 (2013).
- Morrison D. J. & Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 7 189–200 (2016).
- Venkatesh M. et al. Symbiotic Bacterial Metabolites Regulate Gastrointestinal Barrier Function via the Xenobiotic Sensor PXR and Toll-like Receptor 4. Immunity 41 296–310 (2014).
- Honda K. & Littman D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535 75–84 (2016).
- Furusawa Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504 446–450 (2013).
- Arpaia N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504 451 (2013).
- MukhopadhyaI. & Louis P. Gut microbiota-derived short-chain fatty acids and their role in human health and disease. Nat Rev Microbiol 23 635–651 (2025).
- Singh V. et al. Butyrate producers “The Sentinel of Gut”: Their intestinal significance with and beyond butyrate and prospective use as microbial therapeutics. Front.Microbiol. 13 (2023).
- Ren T. et al. Indole Propionic Acid Regulates Gut Immunity: Mechanisms of Metabolite-Driven Immunomodulation and Barrier Integrity. JMicrobiolBiotechnol 35 e2503045 (2025).
- Cryan J. F. et al. The Microbiota-Gut-Brain Axis. Physiological Reviews 99 1877–2013 (2019).
- Matsuura M. Structural Modifications of Bacterial Lipopolysaccharide that Facilitate Gram-Negative Bacteria Evasion of Host Innate Immunity. Front. Immunol. 4 (2013).
- ZamyatinaA. & Heine H. Lipopolysaccharide Recognition in the Crossroads of TLR4 and Caspase-4/11 Mediated Inflammatory Pathways. Front. Immunol. 11 (2020).
- Zhu M. et al. C-reactive protein and cancer risk: a pan-cancer study. BMC Med 20 301 (2022).
- RidkerP. M. et al. Relationship of C-reactive protein reduction to cardiovascular event reduction following treatment with canakinumab. The Lancet 391 319–328 (2018).
- Pradhan A. D. Manson J. E. Rifai N. Buring J. E. &RidkerP. M. C-Reactive Protein Interleukin 6 and Risk of Developing Type 2 Diabetes Mellitus. JAMA 286 327–334 (2001).
- d’HennezelE. Abubucker S. Murphy L. O. & Cullen T. W. Total Lipopolysaccharide from the Human Gut Microbiome Silences Toll-Like Receptor Signaling. mSystems 2 e00046-17 (2017).
- Khorsand B. et al. Overrepresentation of Enterobacteriaceae and Escherichia coli is the major gut microbiome signature in Crohn’s disease and ulcerative colitis. Front. Cell. Infect.Microbiol. 12 (2022).
- Thompson K. N. et al. Alterations in the gut microbiome implicate key taxa and metabolic pathways across inflammatory arthritis phenotypes. Science Translational Medicine 15 eabn4722 (2023).
- Desai M. S. et al. A Dietary Fiber-Deprived Gut Microbiota Degrades the Colonic Mucus Barrier and Enhances Pathogen Susceptibility. Cell 167 1339-1353.e21 (2016).
- Zheng J. et al.Noninvasivemicrobiome-based diagnosis of inflammatory bowel disease. Nat Med 30 3555–3567 (2024).
- Qin J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490 55 (2012).
- Tuomainen M. et al. Associations of serumindolepropionicacid a gut microbiota metabolite with type 2 diabetes and low-grade inflammation in high-risk individuals. Nutrition & Diabetes 8 35 (2018).
- Peron G. et al. A Polyphenol-Rich Diet Increases the Gut Microbiota Metabolite Indole 3-Propionic Acid in Older Adults with Preserved Kidney Function. Molecular Nutrition & Food Research 66 2100349 (2022).
- SpraggeF. et al. Microbiome diversity protects against pathogens by nutrient blocking. Science 382 eadj3502 (2023).
- ByndlossM. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357 570–575 (2017).