What is the
gut microbiome
The gut microbiome is the community of microorganisms, together with their genes and metabolic functions, that live in the gastrointestinal tract. While microbes exist across multiple body sites, the gut microbiome is the most extensively studied because of its density, diversity and metabolic activity.
To give an idea of scale, the total number of these microorganisms equals or exceeds the total number of human cells in the body. These microbes have evolved with us over hundreds of thousands of years and play an important role in health and disease.
Most of the microbes that make up the gut microbiome are bacteria, although fungi, viruses, and archaea can also be present. While microbes live along the entire digestive tract, the largest concentration live in the large intestine (colon), where food residues are fermented.
The gut microbiome is best understood as an ecosystem: a living community of many different microbes that interact with each other and with the human body. Like ecosystems in nature, it has food webs where some microbes break down complex dietary fibres into smaller compounds that other microbes then use. Because different microbes can often perform similar jobs (known as functional redundancy), the overall functions of the ecosystem may matter more for health than the presence or absence of any single species.2,3
The gut microbiome is
a measurable biological system
The gut microbiome is fundamental to normal human physiology. It is a dynamic ecosystem that influences immune regulation, gut barrier integrity, metabolism, and systemic signalling.⁴
This is a two-way relationship. The microbiome contributes
to metabolic activity, produces bioactive compounds, and participates in immune signalling.
In turn, the body provides nutrients that support gut microbes
and influences how they behave. Diet, immune activity, and gut conditions all shape which microbes thrive and what they do.4,5
The microbiome is an integral biological system with meaningful implications for complex and multi-system presentations. Clinically, this is especially important for patients whose symptoms don’t fit neatly into a single diagnosis or one clear cause.
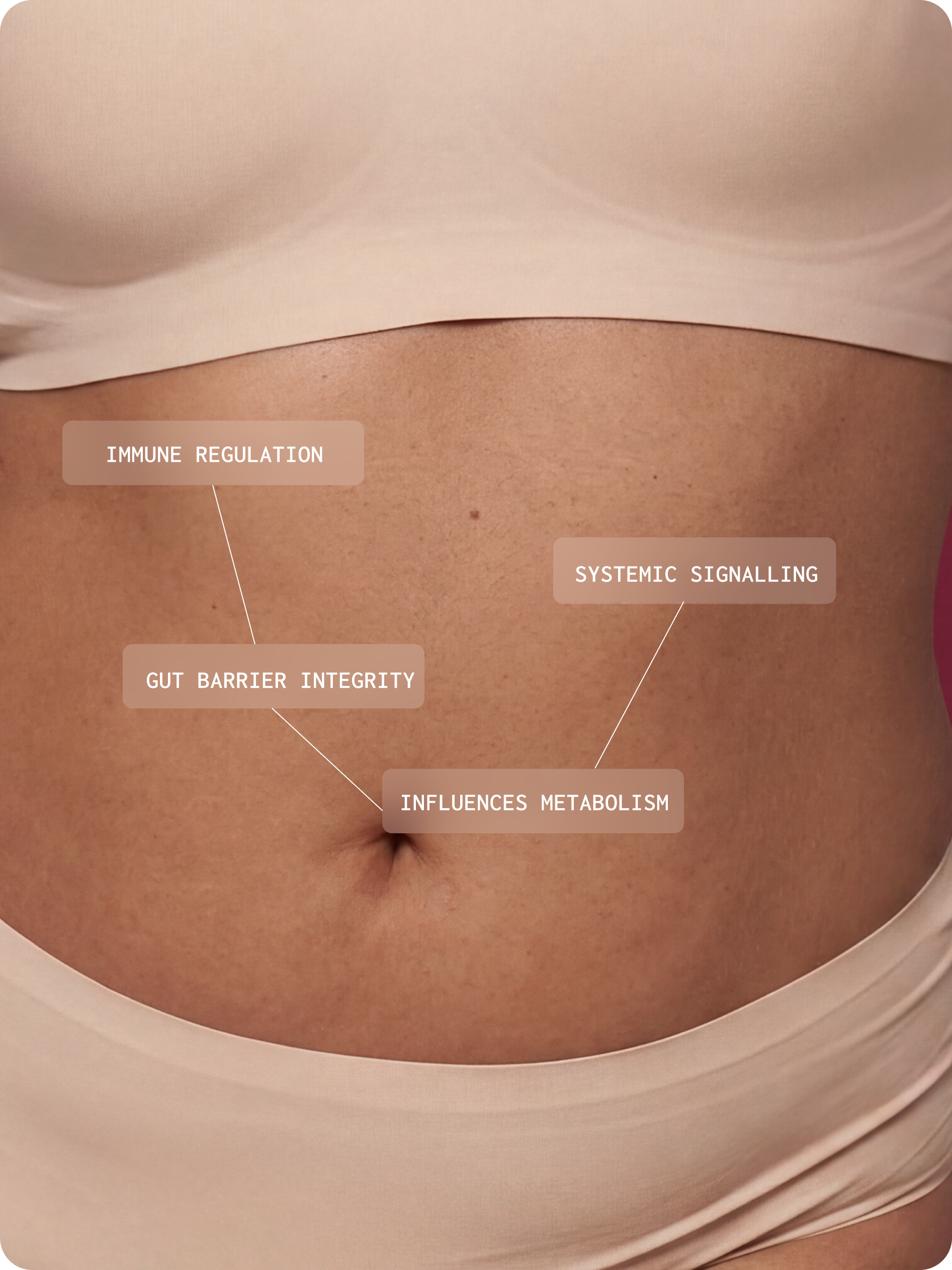
The gut microbiome is composed of diverse microbial communities
The gut microbiome is a diverse community of microorganisms that actively shape gut function through metabolism, competition, and interactions with the immune system and gut lining. This community includes:
Bacteria
Archaea
Viruses
Fungi
The healthy function of the microbiome depends on a balanced ecosystem
The role a microbe plays is not fixed, it depends on the gut environment and the overall ecological balance of the community. Host-microbe interactions generally fall into four broad categories: commensal, mutualistic, opportunistic and pathogenic. These categories are not rigid. The same organism can shift from mutualistic or commensal to opportunistic depending on changes in diet, immune status, or the surrounding microbial community.⁴
COMMENSAL
Present without clearly measurable benefit or harm to the host -- though as detection methods improve, truly neutral organisms are increasingly rare.
MUTUALISTIC
Contributes positively to metabolism, immune regulation, or barrier integrity.
OPPORTUNISTIC
Typically, harmless but may cause harm when ecological balance is disrupted, for example, after antibiotic use or during immune suppression.
PHATHOGENIC
Capable of causing disease or disrupting normal function.
Microbes convert fuel into compounds that influence health
Gut microbes feed on available substrates, including dietary fibre, proteins, and host-derived compounds, and convert them into metabolites that interact with body systems.4,6 Some of these metabolites act locally within the gut, influencing barrier integrity and immune signalling. Others enter the bloodstream and affect organs and processes throughout the body.
What is produced depends on:
Species: The microbes present
Functional pathways: The key metabolic functions they perform
Dominant substrates: The main fuel sources they utilise
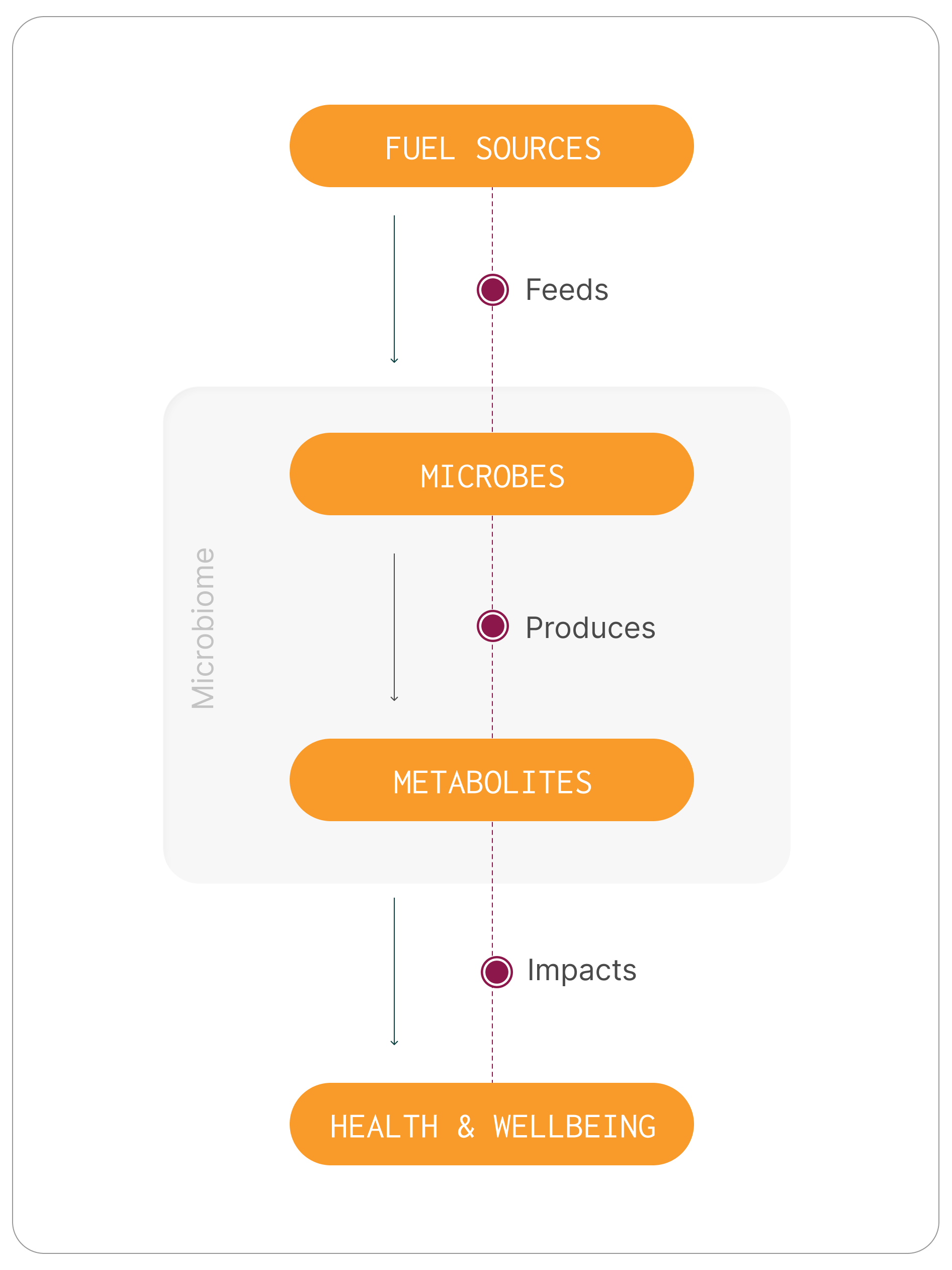
This is why microbiome assessment increasingly focuses on functional capacity and metabolic output — not just species detection.
Knowing which microbes are present isn’t enough, you need to know what the ecosystem is doing.
Microbiome composition varies significantly between individuals. ⁷ While certain microbial groups are commonly found across populations, the relative abundance, diversity, and functional output differ from person to person.
This variability is shaped by many factors – including diet, medication use, genetics, environmental exposures, and lifestyle, all of which influence both microbial composition and metabolic capacity. ⁷
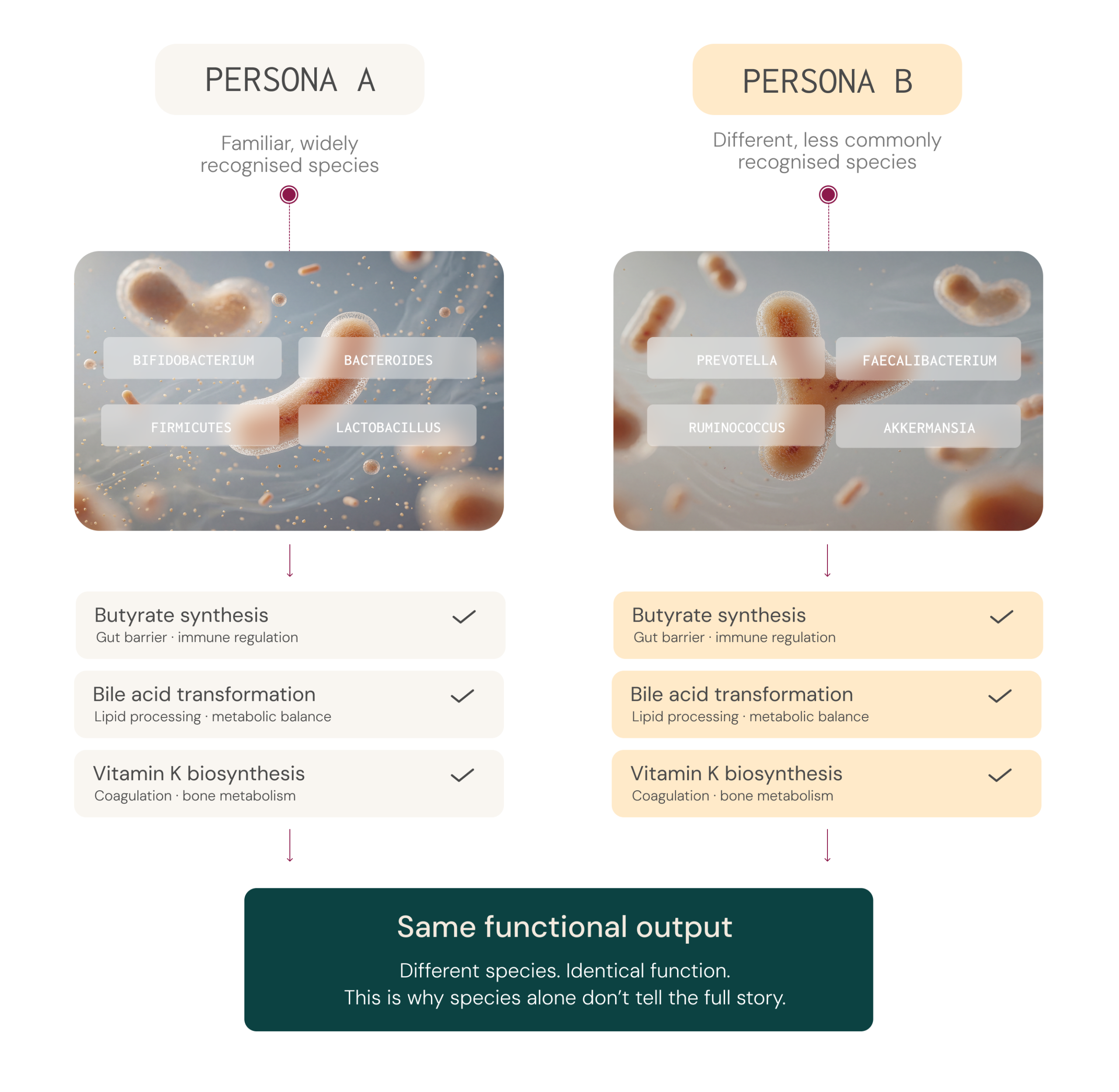
Two individuals can share similar broad microbial groups yet differ considerably in their specific species or strains,² and those species and strains may carry different genes with different metabolic capacities.8,9 The reverse is also true: two people with very different microbial profiles may share comparable functional outputs, because different species can produce the same metabolites.2,3,10
This is why microbiome assessment increasingly focuses on functional capacity and metabolic output – such as butyrate production or mucin degradation, rather than species detection alone.
The microbiome influences health through four key pathways
The gut microbiome influences health not by controlling organs directly, but through the metabolites it produces and the signals they generate.
While the microbiome's influence on the body is broad, four well-characterised functions are particularly relevant to clinical practice: metabolism, immune regulation, gut barrier integrity, and systemic signalling. Importantly, this relationship is bidirectional - the body's environment, including diet, immune activity, and gut conditions, also shapes the microbiome.
Metabolism
Microbial metabolites influence energy use, lipid processing and systemic metabolic balance.
Gut microbes act like a metabolic organ, producing thousands of different metabolites from dietary and host-derived compounds.11,12 Some of these metabolites stay in the gut, where they can affect inflammation, transit time, and the gut lining. Others cross into the bloodstream and can influence inflammation, energy balance, lipid processing, and metabolic regulation throughout the body.11,12
Immune regulation
Continuous exposure to microbial signals helps calibrate immune tolerance and responsiveness.
The microbiome continuously interacts with immune cells within the gut-associated lymphoid tissue (GALT).14 Microbial antigens and metabolites help calibrate immune responsiveness – supporting tolerance to harmless signals while maintaining the ability to respond to genuine threats. For example, some short-chain fatty acids promote the production of regulatory T cells that help prevent over-reactive inflammation.15,16 Rather than simply stimulating immunity, the microbiome plays a central role in immune homeostasis.14
Gut barrier function
Microbial activity supports epithelial integrity and regulated immune signalling.
The gut barrier is a set of layers that separates gut contents from the bloodstream: a mucus layer, a single layer of epithelial cells, and tight junctions between those cells. Microbial metabolites – including butyrate and some tryptophan-derived indoles, can strengthen this barrier by providing an energy source to gut cells and promoting tight junction integrity.6,13,17-19 Thus, barrier function is not purely structural, it is actively maintained by microbial metabolism.
Systemic signalling
Microbial compounds contribute to neuroimmune signalling involved in stress, inflammation, and metabolic integration.
Microbial metabolites interact with the body’s receptors and influence signalling pathways beyond the gut, including enteroendocrine signalling, neural circuits, and immune pathways.4,20 The gut and brain communicate through nerves (including the vagus nerve), hormones, and immune signals. Gut microbes can influence these pathways by producing or modifying neuroactive compounds and by modulating inflammation levels.20
The microbiome functions as part of human biology — not separate from it.
What shapes a person's gut microbiome
The gut microbiome is not static, it is continually adjusting to its environment. A range of modifiable and non-modifiable factors influence which microbes thrive and what they do, many of which are modifiable.⁷
Diet
Medications
Infections and inflammation
Lifestyle and environment
What happens when the microbiome is out of balance
There is no single "perfect" microbiome. Healthy people can have very different microbial profiles. However, when the balance shifts significantly, it can affect what the microbiome produces and how the body responds. An imbalanced microbiome — sometimes called dysbiosis — often involves one or more of the following:
Loss of helpful microbes (for example, fibre-fermenting short-chain fatty acid producers)
Overgrowth of potentially harmful microbes (sometimes called pathobionts)
Reduced diversity (fewer different types of microbes)
A shift in what the microbiome is doing — such as producing fewer protective molecules and more inflammatory molecules
How imbalances can affect the immune system and inflammation

Pro-inflammatory LPS (endotoxin)
Specific types of LPS (hexa-acylated or higher) are strong triggers for inflammation because they can activate a receptor on immune cells called TLR4, which can initiate an inflammatory cascade. Other types of LPS are weaker and may even help block this signalling. When the balance shifts toward bacteria that produce strongly inflammatory LPS, inflammatory signalling can increase. Increased levels of these bacteria have been observed in Crohn’s disease, rheumatoid arthritis, and ankylosing spondylitis.
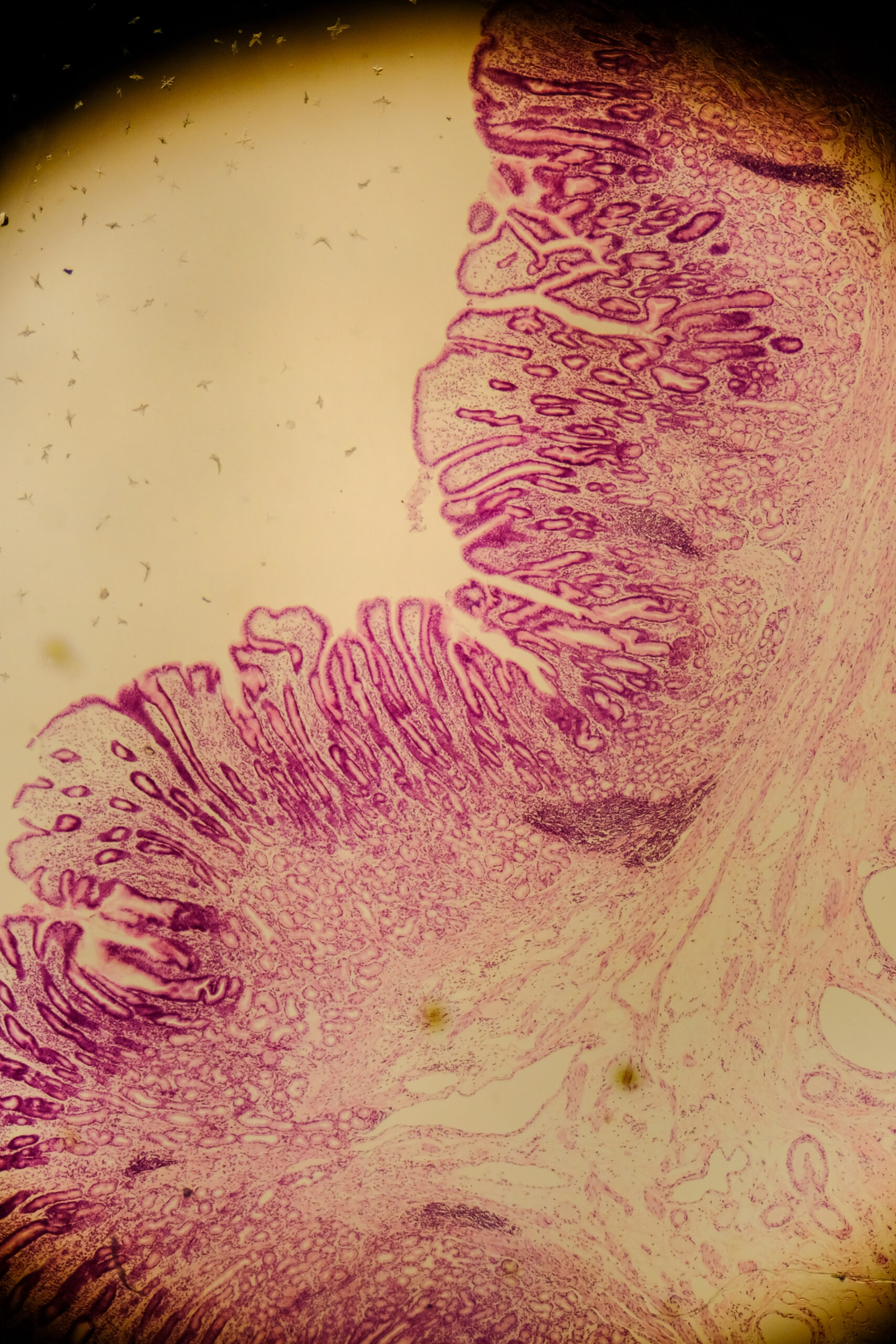
Mucus layer damage when fibre is low
If dietary fibre is limited, some microbes can switch to using mucus as a fuel source. In animal studies, a fibre-deprived diet increased mucus-degrading bacteria, thinned the mucus barrier, and increased susceptibility to infection and inflammation. In humans, the relative abundance of mucin-degrading pathways has been positively correlated with faecal calprotectin, a clinical marker of inflammation in the gut.
Lower butyrate and IPA-producing species
Butyrate is a short-chain fatty acid that serves as the primary energy source for colon cells and plays a key role in immune regulation. Indole-propionic acid (IPA) is a tryptophan-derived metabolite also involved in immune regulation and maintaining intestinal barrier integrity. Reduced levels of butyrate-producing species and plasma IPA have been associated with inflammatory bowel diseases and type 2 diabetes.
Defending against pathogens
A diverse microbiome with adequate levels of butyrate producers can help prevent overgrowth of harmful microbes by competing for space and nutrients, producing antimicrobial substances, and maintaining key signalling pathways. When the microbiome is disrupted, for example after antibiotics, this protective function can be compromised, leaving people more susceptible to pathogen infections.
What matters most in clinical interpretation
Because the microbiome interacts with multiple biological systems, it can provide additional interpretive context in chronic and complex presentations, particularly when traditional markers do not fully explain a patient's symptoms.
The microbiome is not best understood through a "good vs. bad bugs" lens. As outlined in the preceding sections, clinical interpretation is more informative when it considers these key factors: the ecological balance of the community, the functional pathways present, the metabolic output of those pathways, and the individual patient's context.
Ecological balance: community stability and diversity
Functional pathways: what microbes do, not just which are present
Metabolic output: the compounds produced and their effects
Patient context: immune status, diet, and environment
Key Takeaway
The gut microbiome is a measurable, modifiable part of human physiology. It functions as an ecosystem — and like any ecosystem, its health depends on the balance, diversity, and functional capacity of the community as a whole, not just the presence or absence of individual organisms.
For clinicians, the most useful insight from the microbiome is often functional rather than taxonomic: what the microbial community is producing, how that interacts with the patient’s immune system, gut barrier, and metabolic regulation, and how those outputs may be shifting in the context of that individual’s diet, medications, and clinical picture.
For clinicians, microbiome insights provide an additional lens for understanding systemic regulation, particularly in chronic and complex symptom patterns.
As our understanding of the microbiome deepens, so does the opportunity to use it as a meaningful part of clinical assessment, particularly for patients with chronic or multi-system presentations where traditional markers tell only part of the story.
Fundamentals webinar
If you’d like to join our live event to dive deeper into this register for our Fundamentals webinar.
References:
- Sender R. Fuchs S. & Milo R. Revised Estimates for the Number of Human and Bacteria Cells in the Body.PLoSBiology 14 e1002533 (2016).
- HuttenhowerC. et al. Structure function and diversity of the healthy human microbiome. Nature 486 207–214 (2012).
- Jiang Y. Che L. & Li S. C. Deciphering the personalized functional redundancy hierarchy in the gut microbiome. Microbiome 14 17 (2025).
- Fan Y. & Pedersen O. Gut microbiota in human metabolic health and disease. Nat. Rev.Microbiol. 19 55–71 (2021).
- Lynch S. V. & Pedersen O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375 2369–2379 (2016).
- Koh A. DeVadderF. Kovatcheva-Datchary P. & Bäckhed F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 165 1332–1345 (2016).
- ZhernakovaA. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352 565–569 (2016).
- De Filippis F. et al. Distinct genetic and functional traits of human intestinalPrevotellacopri strains are associated with different habitual diets. Cell Host Microbe 25 444–453 (2019).
- Zhang Z. J. et al.Comprehensive analyses of a large human gutBacteroidales culture collection reveal species- and strain-level diversity and evolution. Cell Host Microbe 32 1853–1867 (2024).
- Tian L. et al. Deciphering functional redundancy in the human microbiome. Nat Commun 11 6217 (2020).
- denBestenG. et al. The role of short-chain fatty acids in the interplay between diet gut microbiota and host energy metabolism. Journal of Lipid Research 54 2325–2340 (2013).
- Morrison D. J. & Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 7 189–200 (2016).
- Venkatesh M. et al. Symbiotic Bacterial Metabolites Regulate Gastrointestinal Barrier Function via the Xenobiotic Sensor PXR and Toll-like Receptor 4. Immunity 41 296–310 (2014).
- Honda K. & Littman D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535 75–84 (2016).
- Furusawa Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504 446–450 (2013).
- Arpaia N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504 451 (2013).
- MukhopadhyaI. & Louis P. Gut microbiota-derived short-chain fatty acids and their role in human health and disease. Nat Rev Microbiol 23 635–651 (2025).
- Singh V. et al. Butyrate producers “The Sentinel of Gut”: Their intestinal significance with and beyond butyrate and prospective use as microbial therapeutics. Front.Microbiol. 13 (2023).
- Ren T. et al. Indole Propionic Acid Regulates Gut Immunity: Mechanisms of Metabolite-Driven Immunomodulation and Barrier Integrity. JMicrobiolBiotechnol 35 e2503045 (2025).
- Cryan J. F. et al. The Microbiota-Gut-Brain Axis. Physiological Reviews 99 1877–2013 (2019).
- Matsuura M. Structural Modifications of Bacterial Lipopolysaccharide that Facilitate Gram-Negative Bacteria Evasion of Host Innate Immunity. Front. Immunol. 4 (2013).
- ZamyatinaA. & Heine H. Lipopolysaccharide Recognition in the Crossroads of TLR4 and Caspase-4/11 Mediated Inflammatory Pathways. Front. Immunol. 11 (2020).
- Zhu M. et al. C-reactive protein and cancer risk: a pan-cancer study. BMC Med 20 301 (2022).
- RidkerP. M. et al. Relationship of C-reactive protein reduction to cardiovascular event reduction following treatment with canakinumab. The Lancet 391 319–328 (2018).
- Pradhan A. D. Manson J. E. Rifai N. Buring J. E. &RidkerP. M. C-Reactive Protein Interleukin 6 and Risk of Developing Type 2 Diabetes Mellitus. JAMA 286 327–334 (2001).
- d’HennezelE. Abubucker S. Murphy L. O. & Cullen T. W. Total Lipopolysaccharide from the Human Gut Microbiome Silences Toll-Like Receptor Signaling. mSystems 2 e00046-17 (2017).
- Khorsand B. et al. Overrepresentation of Enterobacteriaceae and Escherichia coli is the major gut microbiome signature in Crohn’s disease and ulcerative colitis. Front. Cell. Infect.Microbiol. 12 (2022).
- Thompson K. N. et al. Alterations in the gut microbiome implicate key taxa and metabolic pathways across inflammatory arthritis phenotypes. Science Translational Medicine 15 eabn4722 (2023).
- Desai M. S. et al. A Dietary Fiber-Deprived Gut Microbiota Degrades the Colonic Mucus Barrier and Enhances Pathogen Susceptibility. Cell 167 1339-1353.e21 (2016).
- Zheng J. et al.Noninvasivemicrobiome-based diagnosis of inflammatory bowel disease. Nat Med 30 3555–3567 (2024).
- Qin J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490 55 (2012).
- Tuomainen M. et al. Associations of serumindolepropionicacid a gut microbiota metabolite with type 2 diabetes and low-grade inflammation in high-risk individuals. Nutrition & Diabetes 8 35 (2018).
- Peron G. et al. A Polyphenol-Rich Diet Increases the Gut Microbiota Metabolite Indole 3-Propionic Acid in Older Adults with Preserved Kidney Function. Molecular Nutrition & Food Research 66 2100349 (2022).
- SpraggeF. et al. Microbiome diversity protects against pathogens by nutrient blocking. Science 382 eadj3502 (2023).
- ByndlossM. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357 570–575 (2017).